Abstract
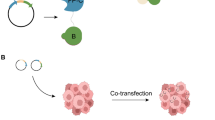
Deciphering protein–protein interactions (PPIs) in vivo is crucial to understand protein function. Bimolecular fluorescence complementation (BiFC) makes applicable the analysis of PPIs in many different native contexts, including human live cells. It relies on the property of monomeric fluorescent proteins to be reconstituted from two separate subfragments upon spatial proximity. Candidate partners fused to such complementary subfragments can form a fluorescent protein complex upon interaction, allowing visualization of weak and transient PPIs. It can also be applied for investigation of distinct PPIs at the same time using a multicolor setup. In this chapter, we provide a detailed protocol for analyzing PPIs by doing BiFC in cultured cells. Proof-of-principle experiments rely on the complementation property between the N-terminal fragment of mVenus (designated VN173) and the C-terminal fragment of mCerulean (designated CC155) and the partnership between HOXA7 and PBX1 proteins. This protocol is compatible with any other fluorescent complementation pair fragments and any type of candidate interacting proteins.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Kerppola TK (2006) Visualization of molecular interactions by fluorescence complementation. Nat Rev Mol Cell Biol 7:449–456
Hu C-D, Kerppola TK (2003) Simultaneous visualization of multiple protein interactions in living cells using multicolor fluorescence complementation analysis. Nat Biotechnol 21:539–545
Kerppola TK (2006) Design and implementation of bimolecular fluorescence complementation (BiFC) assays for the visualization of protein interactions in living cells. Nat Protoc 1:1278–1286
Rizzo MA, Springer GH, Granada B et al (2004) An improved cyan fluorescent protein variant useful for FRET. Nat Biotechnol 22:445–449
Nagai T, Ibata K, Park ES et al (2002) A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat Biotechnol 20:87–90
Bhat RA, Lahaye T, Panstruga R (2006) The visible touch: in planta visualization of protein-protein interactions by fluorophore-based methods. Plant Methods 2:12
Hu C-D, Chinenov Y, Kerppola TK (2002) Visualization of interactions among bZIP and Rel family proteins in living cells using bimolecular fluorescence complementation. Mol Cell 9:789–798
Vogel SS, Thaler C, Koushik SV (2006) Fanciful FRET. Sci STKE 2006:re2
Hu C-D, Grinberg AV, Kerppola TK (2006) Visualization of protein interactions in living cells using bimolecular fluorescence complementation (BiFC) analysis. Curr Protoc Cell Biol Chapter 21:Unit 21.3
Magli MC, Largman C, Lawrence HJ (1997) Effects of HOX homeobox genes in blood cell differentiation. J Cell Physiol 173:168–177
Moens CB, Selleri L (2006) Hox cofactors in vertebrate development. Dev Biol 291:193–206
Passner JM, Ryoo HD, Shen L et al (1999) Structure of a DNA-bound Ultrabithorax-Extradenticle homeodomain complex. Nature 397:714–719
Dard A, Reboulet J, Jia Y et al (2018) Human HOX proteins use diverse and context-dependent motifs to interact with TALE class cofactors. Cell Rep 22:3058–3071
LaRonde-LeBlanc NA, Wolberger C (2003) Structure of HoxA9 and Pbx1 bound to DNA: Hox hexapeptide and DNA recognition anterior to posterior. Genes Dev 17:2060–2072
Shyu YJ, Liu H, Deng X et al (2006) Identification of new fluorescent protein fragments for bimolecular fluorescence complementation analysis under physiological conditions. BioTechniques 40:61–66
Shaner NC, Campbell RE, Steinbach PA et al (2004) Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat Biotechnol 22:1567–1572
Schindelin J, Arganda-Carreras I, Frise E et al (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9:676–682
Blogger G. When is a monomer not a monomer? The top three ways your favorite fluorescent protein oligomerizes in cells. https://blog.addgene.org/when-is-a-monomer-not-a-monomer-the-top-three-ways-your-favorite-fluorescent-protein-oligomerizes-in-cells
Zacharias DA, Violin JD, Newton AC et al (2002) Partitioning of lipid-modified monomeric GFPs into membrane microdomains of live cells. Science 296:913–916
Kudla J, Bock R (2016) Lighting the way to protein-protein interactions: recommendations on best practices for bimolecular fluorescence complementation analyses. Plant Cell 28:1002–1008
Horstman A, Tonaco IAN, Boutilier K et al (2014) A cautionary note on the use of split-YFP/BiFC in plant protein-protein interaction studies. Int J Mol Sci 15:9628–9643
Xia J, Kong L, Zhou L-J et al (2018) Genome-wide bimolecular fluorescence complementation-based proteomic analysis of toxoplasma gondii ROP18’s human Interactome shows its key role in regulation of cell immunity and apoptosis. Front Immunol 9:61
Yue L, Li L, Li D et al (2017) High-throughput screening for Survivin and Borealin interaction inhibitors in hepatocellular carcinoma. Biochem Biophys Res Commun 484:642–647
Lepur A, Kovačević L, Belužić R et al (2016) Combining unique multiplex gateway cloning and bimolecular fluorescence complementation (BiFC) for high-throughput screening of protein–protein interactions. J Biomol Screen 21:1100–1111
Vidi P-A, Przybyla JA, Hu C-D et al (2010) Visualization of G protein-coupled receptor (GPCR) interactions in living cells using bimolecular fluorescence complementation (BiFC). Curr Protoc Neurosci Chapter 5:Unit-5.29
PB helping cells and sections to stick: cleaning, sterilising and coating slides and coverslips | Agar Scientific. http://www.agarscientific.net/helping-cells-and-sections-to-stick-cleaning-sterilising-and-coating-slides-and-coverslips/
Ando K, Parsons MJ, Shah RB et al (2017) NPM1 directs PIDDosome-dependent caspase-2 activation in the nucleolus. J Cell Biol 216:1795–1810
Szymczak AL, Vignali DAA (2005) Development of 2A peptide-based strategies in the design of multicistronic vectors. Expert Opin Biol Ther 5:627–638
Liu Z, Chen O, Wall JBJ et al (2017) Systematic comparison of 2A peptides for cloning multi-genes in a polycistronic vector. Sci Rep 7:2193
Acknowledgments
Research in the laboratory of S. Merabet is supported by Association pour la Recherche sur le Cancer (ARC, PJA20141202007); Fondation pour la Recherche Médicale (FRM, DEQ. 20170336732); Ligue Nationale Contre le Cancer, Centre National de Recherche Scientifique (CNRS); CEFIPRA (5503-2); CNRS; and Ecole Normale Supérieure (ENS) de Lyon. We further thank the China Scholarship Council (CSC, File No. 201708070003) for the doctoral grant to Y. Jia.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Jia, Y., Bleicher, F., Reboulet, J., Merabet, S. (2021). Bimolecular Fluorescence Complementation (BiFC) and Multiplexed Imaging of Protein–Protein Interactions in Human Living Cells. In: Zamir, E. (eds) Multiplexed Imaging. Methods in Molecular Biology, vol 2350. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1593-5_12
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1593-5_12
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1592-8
Online ISBN: 978-1-0716-1593-5
eBook Packages: Springer Protocols