Abstract
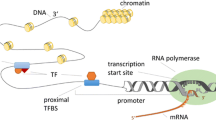
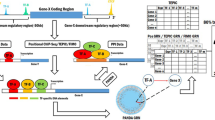
The cell expresses various genes in specific contexts with respect to internal and external perturbations to invoke appropriate responses. Transcription factors (TFs) orchestrate and define the expression level of genes by binding to their regulatory regions. Dysregulated expression of TFs often leads to aberrant expression changes of their target genes and is responsible for several diseases including cancers. In the last two decades, several studies experimentally identified target genes of several TFs. However, these studies are limited to a small fraction of the total TFs encoded by an organism, and only for those amenable to experimental settings. Experimental limitations lead to many computational techniques having been proposed to predict target genes of TFs. Linear modeling of gene expression is one of the most promising computational approaches, readily applicable to the thousands of expression datasets available in the public domain across diverse phenotypes. Linear models assume that the expression of a gene is the sum of expression of TFs regulating it. In this chapter, I introduce mathematical programming for the linear modeling of gene expression, which has certain advantages over the conventional statistical modeling approaches. It is fast, scalable to genome level and most importantly, allows mixed integer programming to tune the model outcome with prior knowledge on gene regulation.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
CRICK FH (1958) On protein synthesis. Symp Soc Exp Biol 12:138–163. http://www.ncbi.nlm.nih.gov/pubmed/13580867
Muley VY, Pathania A (2017) Gene Expression. In: Vonk J, Shackelford T (eds) Encycl Anim Cogn Behav. Springer, Cham
Pathania A, Muley VY (2017) Gene expression profiling. In: Vonk J, Shackelford T (eds) Encycl Anim Cogn Behav. Springer, Cham
Law CW, Alhamdoosh M, Su S et al (2018) RNA-seq analysis is easy as 1-2-3 with limma, Glimma and edgeR. F1000 Res 5:1408. https://doi.org/10.12688/f1000research.9005.3
Cheng C, Alexander R, Min R, Leng J, Yip KY, Rozowsky J et al (2012) Understanding transcriptional regulation by integrative analysis of TF binding data. Genome Res 22(9):1658–1667
Taylor RC, Acquaah-Mensah G, Singhal M, Malhotra D, Biswal S (2008) Network inference algorithms elucidate Nrf2 regulation of mouse lung oxidative stress. PLoS Comput Biol 4(8):e1000166
Setty M, Helmy K, Khan AA, Silber J, Arvey A, Neezen F et al (2012) Inferring transcriptional and microRNA-mediated regulatory programs in glioblastoma. Mol Syst Biol 8:605
Poos AM, Maicher A, Dieckmann AK, Oswald M, Eils R, Kupiec M et al (2016) Mixed Integer Linear Programming based machine learning approach identifies regulators of telomerase in yeast. Nucleic Acids Res 44:e93. https://doi.org/10.1093/nar/gkw111
Marbach D, Costello JC, Küffner R, Vega NM, Prill RJ, Camacho DM et al (2012) Wisdom of crowds for robust gene network inference. Nat Methods 9:796–804
Gurobi Optimization LLC. Gurobi optimizer reference manual. 2020. http://www.gurobi.com
Schacht T, Oswald M, Eils R, Eichmüller SB, König R (2014) Estimating the activity of TFs by the effect on their target genes. Bioinformatics 30:i401–i407. https://doi.org/10.1093/bioinformatics/btu446
Vaquerizas JM, Kummerfeld SK, Teichmann SA, Luscombe NM (2009) A census of human TFs: Function, expression and evolution. Nat Rev Genet 10:252–263. http://www.ncbi.nlm.nih.gov/pubmed/19274049
Muley VY, López-Victorio CJ, Ayala-Sumuano JT, González-Gallardo A, González-Santos L, Lozano-Flores C et al (2020) Conserved and divergent expression dynamics during early patterning of the telencephalon in mouse and chick embryos. Prog Neurobiol 186:101735
Kernohan KD, Jiang Y, Tremblay DC, Bonvissuto AC, Eubanks JH, Mann MRW et al (2010) ATRX Partners with Cohesin and MeCP2 and Contributes to Developmental Silencing of Imprinted Genes in the Brain. Dev Cell 18:191–202. https://linkinghub.elsevier.com/retrieve/pii/S153458071000016X
Fujii Y, Yoshihashi K, Suzuki H, Tsutsumi S, Mutoh H, Maeda S et al (2012) CDX1 confers intestinal phenotype on gastric epithelial cells via induction of stemness-associated reprogramming factors SALL4 and KLF5. Proc Natl Acad Sci U S A 109:20584–20589. http://www.ncbi.nlm.nih.gov/pubmed/23112162
Ostapcuk V, Mohn F, Carl SH, Basters A, Hess D, Iesmantavicius V et al (2018) Activity-dependent neuroprotective protein recruits HP1 and CHD4 to control lineage-specifying genes. Nature 557:739–743. http://www.nature.com/articles/s41586-018-0153-8
Gerstein MB, Kundaje A, Hariharan M, Landt SG, Yan KK, Cheng C et al (2012) Architecture of the human regulatory network derived from ENCODE data. Nature 489:91–100
Acknowledgments
This work was supported by DGAPA-UNAM grant IA203920 to V.Y.M. The author would like to sincerely thank Anne Hahn (Queensland Brain Institute, Australia) for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Muley, V.Y. (2021). Mathematical Programming for Modeling Expression of a Gene Using Gurobi Optimizer to Identify Its Transcriptional Regulators. In: MUKHTAR, S. (eds) Modeling Transcriptional Regulation. Methods in Molecular Biology, vol 2328. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1534-8_6
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1534-8_6
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1533-1
Online ISBN: 978-1-0716-1534-8
eBook Packages: Springer Protocols