Abstract
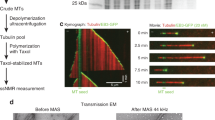
In this chapter, we describe the preparatory and spectroscopic procedures for conducting solid-state NMR experiments on microtubules (MTs) obtained from human cells and their complexes with microtubule-associated proteins (MAPs). Next to labeling and functional assembly of MTs and MT-MAP complexes, we discuss solid-state NMR approaches, including fast MAS and hyperpolarization methods that can be used to examine these systems. Such studies can provide novel insight into the dynamic properties of MTs and MT-MAP complexes.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Renault M, Cukkemane A, Baldus M (2010) Solid-state NMR spectroscopy on complex biomolecules. Angew Chem Int Ed 49:8346–8357
Quinn CM, Polenova T (2017) Structural biology of supramolecular assemblies by magic-angle spinning NMR spectroscopy. Q Rev Biophys 50:e1
Boisbouvier J, Kay LE (2018) Advanced isotopic labeling for the NMR investigation of challenging proteins and nucleic acids. J Biomol NMR 71:115–117
Kaplan M, Narasimhan S, de Heus C et al (2016) EGFR dynamics change during activation in native membranes as revealed by NMR. Cell 167:1241–1251
Narasimhan S, Scherpe S, Paioni A et al (2019) DNP supported solid-state NMR of proteins inside mammalian cells. Angew Chem Int Ed Eng 58:12969–12973
Theillet F-X, Binolfi A, Bekei B et al (2016) Structural disorder of monomeric α-synuclein persists in mammalian cells. Nature 530:45–50
Kumar A, Blommers MJ, Krastel P et al (2010) Interaction of Epothilone B (Patupilone) with microtubules as detected by two-dimensional solid-state NMR spectroscopy. Angew Chem Int Ed Eng 49:7504–7507
Yan S, Guo C, Hou G et al (2015) Atomic-resolution structure of the CAP-Gly domain of dynactin on polymeric microtubules determined by magic angle spinning NMR spectroscopy. Proc Natl Acad Sci U S A 112:14611–14616
Atherton J, Jiang K, Stangier MM et al (2017) A structural model for microtubule minus-end recognition and protection by CAMSAP proteins. Nat Struct Mol Biol 24:931–943
Kubo S, Nishida N, Udagawa Y et al (2013) A gel-encapsulated bioreactor system for NMR studies of protein–protein interactions in living mammalian cells. Angew Chem Int Ed Eng 52:1208–1211
Kesten C, Wallmann A, Schneider R et al (2019) The companion of cellulose synthase 1 confers salt tolerance through a Tau-like mechanism in plants. Nat Commun 10:857
Kadavath H, Fontela YC, Jaremko M et al (2018) The binding mode of a tau peptide with tubulin. Angew Chem Int Ed Eng 57:3246–3250
Atherton J, Luo Y, Xiang S et al (2019) Structural determinants of microtubule minus end preference in CAMSAP CKK domains. Nat Commun 10:5236
Luo Y, Xaing S, Hooikaas PJ et al (2020) Direct observation of dynamic protein interactions involving human microtubules using solid-state NMR spectroscopy. Nat Commun 11:18
Damman R, Schutz S, Luo Y et al (2019) Atomic-level insight into mRNA processing bodies by combining solid and solution-state NMR spectroscopy. Nat Commun 10:4536
Souphron, J, Bodakuntla, S, Jijumon, AS et al (2019) Purification of tubulin with controlled post-translational modifications by polymerization–depolymerization cycles Nature Protocols, 14, 1634–1660
Fung B, Khitrin AK, Ermolaev K (2000) An improved broadband decoupling sequence for liquid crystals and solids. J Magn Reson 142:97–101
Bennett AE, Rienstra CM, Griffiths J (1998) Homonuclear radio frequency-driven recoupling in rotating solids. J Chem Phys 108:9463–9479
Sauvée C, Rosay M, Casano G et al (2013) Highly efficient, water-soluble polarizing agents for dynamic nuclear polarization at high frequency. Angew Chem Int Ed Eng 52:10858–10861
Gradmann S, Ader C, Heinrich I et al (2012) Rapid prediction of multi-dimensional NMR data sets. J Biomol NMR 54:377–387
Acknowledgments
We thank Gert Folkers for helpful discussions and Johan van der Zwan for excellent technical support. This work was supported by the Dutch Science Foundation NWO (VENI grant 722.016.002 to SX, NWO-Groot grant 175.010.2009.002, and TOP-PUNT grant to MB) and by uNMR-NL, the National Roadmap Large-Scale NMR Facility of the Netherlands (grant 184.032.207). C.J. is supported by the grants ANR-10-IDEX-0001-02 PSL, ANR-11-LBX-0038, FRM DEQ20170336756. S.B. was supported by the FRM grant FDT201805005465, and CEFIPRA 5703-1, and J.A.S. by the European Union’s Horizon 2020 Marie Skłodowska-Curie grant agreement No. 675737, and the FRM grant FDT201904008210.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Luo, Y. et al. (2021). Solid-State NMR Spectroscopy for Studying Microtubules and Microtubule-Associated Proteins. In: Owens, R.J. (eds) Structural Proteomics. Methods in Molecular Biology, vol 2305. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1406-8_10
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1406-8_10
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1405-1
Online ISBN: 978-1-0716-1406-8
eBook Packages: Springer Protocols