Abstract
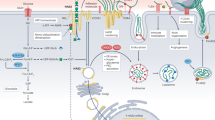
The ubiquitous extracellular glycosaminoglycan hyaluronan (HA) is a polymer composed of repeated disaccharide units of alternating D-glucuronic acid and D-N-acetylglucosamine residues linked via alternating β-1,4 and β-1,3 glycosidic bonds. Emerging data continue to reveal functions attributable to HA in a variety of physiological and pathological contexts. Defining the mechanisms regulating expression of the human hyaluronan synthase (HAS) genes that encode the corresponding HA-synthesizing HAS enzymes is therefore important in the context of HA biology in health and disease. We describe here methods to analyze transcriptional regulation of the HAS and HAS2-antisense RNA 1 genes. Elucidation of mechanisms of HA interaction with receptors such as the cell surface molecule CD44 is also key to understanding HA function. To this end, we provide protocols for fluorescent recovery after photobleaching analysis of CD44 membrane dynamics in the process of fibroblast to myofibroblast differentiation, a phenotypic transition that is common to the pathology of fibrosis of large organs such as the liver and kidney.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Spicer AP, Seldin MF, Olsen AS et al (1997) Chromosomal localization of the human and mouse hyaluronan synthase genes. Genomics 41:493–497
Sayo T, Sugiyama Y, Takahashi Y et al (2002) Hyaluronan synthase 3 regulates hyaluronan synthesis in cultured human keratinocytes. J Investig Dermatol 118:43–48
Monslow J, Williams JD, Norton N et al (2003) The human hyaluronan synthase genes: genomic structures, proximal promoters and polymorphic microsatellite markers. Int J Biochem Cell Biol 35:1272–1283
Chao H, Spicer AP (2005) Natural antisense mRNAs to hyaluronan synthase 2 inhibit hyaluronan biosynthesis and cell proliferation. J Biol Chem 280:27513–27522
Dobzhansky T (1964) Biology, molecular and organismic. Am Zool 4:443–452
Weigel PH, DeAngelis PL (2007) Hyaluronan synthases: a decade-plus of novel glycosyltransferases. J Biol Chem 282:36777–36781
Weigel PH, Hascall VC, Tammi M (1997) Hyaluronan synthases. J Biol Chem 272:13997–14000
Itano N, Sawai T, Yoshida M et al (1999) Three isoforms of mammalian hyaluronan synthases have distinct enzymatic properties. J Biol Chem 274:25085–25092
Rilla K, Oikari S, Jokela TA et al (2013) Hyaluronan synthase 1 (HAS1) requires higher cellular UDP-GlcNAc concentration than HAS2 and HAS3. J Biol Chem 288:5973–5983
Bart G, Ortega-Vico N, Hasinen A et al (2015) Fluorescence resonance energy transfer (FRET) and proximity ligation assays reveal functionally relevant homo- and heteromeric complexes among hyaluronan synthases HAS1, HAS2, and HAS3. J Biol Chem 290(18):11479–11490
Michael DR, Phillips AO, Krupa A et al (2011) The human hyaluronan synthase 2 (HAS2) gene and its natural antisense RNA exhibit coordinated expression in the renal proximal tubular epithelial cell. J Biol Chem 286:19523–19532
Tammi RH, Passi AG, Rilla K (2011) Transcriptional and post-translational regulation of hyaluronan synthesis. FEBS J 278:1419–1428
Vigetti D, Viola M, Karousou E et al (2013) Metabolic control of hyaluronan synthases. Matrix Biol 35:8–13. https://doi.org/10.1016/j.matbio.2013.10.002
Caon I, Parnigoni A, Viola M et al (2020) Cell energy metabolism and hyaluronan synthesis. J Histochem Cytochem 69(1):35–47. https://doi.org/10.1369/0022155420929772
Melero-Fernandez de Mara RM, Arasu UT, Kärnä R et al (2019) Effects of mutations in the post-translational modification sites on the trafficking of hyaluronan synthase 2 (HAS2). Matrix Biol 80:85–103
Yamada Y, Itano N, Hata K-I et al (2004) Differential regulation by IL-1beta and EGF of expression of three different hyaluronan synthases in oral mucosal epithelial cells and fibroblasts and dermal fibroblasts: quantitative analysis using real-time RT-PCR. J Investig Dermatol 122:631–639
Monslow J, Williams JD, Guy CA et al (2004) Identification and analysis of the promoter region of the human hyaluronan synthase 2 gene. J Biol Chem 279:20576–20581
Monslow J, Williams JD, Fraser DJ et al (2006) Sp1 and Sp3 mediate constitutive transcription of the human hyaluronan synthase 2 gene. J Biol Chem 281:18043–18050
Chen L, Neville RD, Michael DR et al (2012) Identification and analysis of the human hyaluronan synthase 1 gene promoter reveals Smad3- and Sp3-mediated transcriptional induction. Matrix Biol 31:373–379
Saavalainen K, Tammi MI, Bowen T et al (2007) Integration of the activation of the human hyaluronan synthase 2 gene promoter by common co-factors of the transcription factors RAR and NF-κB. J Biol Chem 282:11530–11539
Vigetti D, Deleonibus S, Moretto P et al (2014) Natural antisense transcript for Hyaluronan synthase 2 (HAS2-AS1) induces transcription of HAS2 via protein O-GlcNAcylation. J Biol Chem 289(4):28816–28826
Siiskonen H, Oikari S, Pasonen-Seppänen S et al (2015) Hyaluronan synthase 1: a mysterious enzyme with unexpected functions. Front Immunol 6:43
Cho S, Roh K, Park J et al (2017) Hydrolysis of hyaluronic acid in lymphedematous tissue alleviates fibrogenesis via TH1 cell-mediated cytokine expression. Sci Rep 7:35
Lauer-Fields JL, Malkar NB, Richet G et al (2003) Melanoma cell CD44 interaction with the alpha 1(IV) 1263-1277 region from basement membrane collagen is modulated by ligand glycosylation. J Biol Chem 278:14321–14330
Marrero-Diaz R, Bravo-Cordero JJ, Megías D et al (2009) Polarized MT1-MMP-CD44 interaction and CD44 cleavage during cell retraction reveal an essential role for MT1-MMP in CD44-mediated invasion. Cell Motil Cytoskeleton 66:48–61
Midgley AC, Rogers M, Hallet MB et al (2013) Transforming growth factor-β1 (TGF-β1)-stimulated fibroblast to myofibroblast differentiation is mediated by hyaluronan (HA)-facilitated epidermal growth factor receptor (EGFR) and CD44 co-localization in lipid rafts. J Biol Chem 288:14824–14838
Midgley AC, Duggal L, Jenkins R et al (2015) Hyaluronan regulates bone morphogenetic protein-7-dependent prevention and reversal of myofibroblast phenotype. J Biol Chem 290(18):11218–11234
Midgley AC, Oltean S, Hascall V et al (2017) Nuclear hyaluronidase 2 drives alternative splicing of CD44 pre-mRNA to determine profibrotic or antifibrotic cell phenotype. Sci Signal 10:eaao1822
Meran S, Luo DD, Simpson RM et al (2011) Hyaluronan facilitates transforming growth factor-beta1-dependent proliferation via CD44 and epidermal growth factor receptor interaction. J Biol Chem 286:17618–17630
Simpson RM, Wells A, Thomas DW et al (2010) Aging fibroblasts resist phenotypic maturation because of impaired hyaluronan-dependent CD44/epidermal growth factor receptor signaling. Am J Pathol 176:1215–1228
Webber J, Meran S, Steadman R et al (2009) Hyaluronan orchestrates transforming growth factor-beta1-dependent maintenance of myofibroblast phenotype. J Biol Chem 284:9083–9092
Martin J, Midgley A, Meran S et al (2016) Tumor necrosis factor-stimulated gene 6 (TSG-6)-mediated interactions with the inter-α-inhibitor heavy chain 5 facilitate transforming growth factor β1 (TGFβ1)-dependent fibroblast to myofibroblast differentiation. J Biol Chem 291:13789–13801
Midgley AC, Bowen T, Phillips AO et al (2014) MicroRNA-7 inhibition rescues age-associated loss of epidermmal growth factor receptor and hyaluronan-dependent differentiation in fibroblasts. Aging Cell 13:235–244
Untergasser A, Cutcutache I, Koressaar T et al (2012) Primer3-new capabilities and interfaces. Nucleic Acids Res 40:e115
Koressaar T, Remm M (2007) Enhancements and modifications of primer design program Primer3. Bioinformatics 23:1289–1291
Koressaar T, Lepamets M, Kaplinski L et al (2018) Primer3_masker: integrating masking of template sequence with primer design software. Bioinformatics 34:1937–1938
Bowen T, Williams NM, Norton N et al (2001) Mutation screening of the KCNN3 gene reveals a rare frameshift mutation. Mol Psychiatry 6:259–260
Axelrod D, Koppel DE, Schlessinger J et al (1976) Mobility measurement by analysis of fluorescence photobleaching recovery kinetics. Biophys J 16:1055–1069
Acknowledgments
The authors are supported by funding from Kidney Research UK, Medical Research Council UK, Wales Kidney Research Unit, Wellcome Trust, UCB Pharma and the National Natural Science Foundation of China (NSFC) International Young Scholar Research Fellowship (81850410552).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Midgley, A.C., Bowen, T. (2022). Analysis of Human Hyaluronan Synthase Gene Transcriptional Regulation and Downstream Hyaluronan Cell Surface Receptor Mobility in Myofibroblast Differentiation. In: Balagurunathan, K., Nakato, H., Desai, U., Saijoh, Y. (eds) Glycosaminoglycans. Methods in Molecular Biology, vol 2303. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1398-6_36
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1398-6_36
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1397-9
Online ISBN: 978-1-0716-1398-6
eBook Packages: Springer Protocols