Abstract
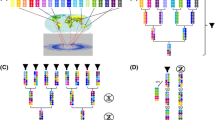
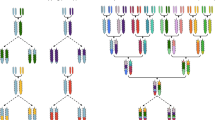
Biparental mapping populations consist of a set of individuals derived from crosses between two parents often belonging to diverse species of a botanical genus and differing in terms of phenotype and traits to share. The development of such recombinant libraries represents a powerful strategy for dissection of the genetic basis of complex traits in crops and these are largely utilized to develop pre-breeding sources to use in crop improvement. This chapter provides an overview of methods and strategies to follow, for the construction of different types of populations, from a plant breeder point of view. Starting from the initial crossing between founder lines toward the further selection steps, here are described the populations commonly established in autogamous species including F2, double haploids, backcrosses and recombinant inbreds, and introgression lines.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
D’Agostino N, Tripodi P (2017) NGS-based genotyping, high-throughput phenotyping and genome-wide association studies laid the foundations for next-generation breeding in horticultural crops. Diversity 9(3):38. https://doi.org/10.3390/d9030038
Moose SP, Mumm RH (2008) Molecular plant breeding as the foundation for 21st century crop improvement. Plant Physiol 147:969–977
Hajjar R, Hodgkin T (2007) The use of wild relatives in crop improvement: a survey of developments over the last 20 years. Euphytica 156:1–13
Nair KP (2019) Gene flow between cultivated plants and their wild relatives. In: Combating global warming. Springer, Cham, pp 49–52. https://doi.org/10.1007/978-3-030-23037-1_10
Xu Y, Li P, Yang Z, Xu C (2017) Genetic mapping of quantitative trait loci in crops. Crop J 5:175–184
Keating BA, Herrero M, Carberry PS, Gardner J, Cole MB (2014) Food wedges: framing the global food demand and supply challenge towards 2050. Glob Food Secur 3:125–132
Xu Y (2010) Molecular plant breeding. CAB International, Wallingford
Schnable PS, Hsia A-P, Nikolau BJ (1998) Genetic recombination in plants. Curr Opin Plant Biol 1(2):123–129. https://doi.org/10.1016/S1369-5266(98)80013-7
Zhang YM, Xu SZ (2004) Mapping quantitative trait loci in F2 incorporating phenotypes of F3 progeny. Genetics 166:1981–1993
Maluszynski M, Kasha KJ, Forster BP, Szarejko I (2003) Doubled haploid production in crop plants: a manual. Kluwer Academic, Dordrecht
Alseekh S, Ofner I, Pleban T, Tripodi P, Di Dato F, Cammareri M, Mohammad A, Grandillo S, Fernie AR, Zamir D (2013) Resolution by recombination: breaking up Solanum pennellii introgressions. Trends Plant Sci 18:536–538
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Tripodi, P. (2021). Methods of Development of Biparental Mapping Populations in Horticultural Crops. In: Tripodi, P. (eds) Crop Breeding. Methods in Molecular Biology, vol 2264. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1201-9_1
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1201-9_1
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1200-2
Online ISBN: 978-1-0716-1201-9
eBook Packages: Springer Protocols