Abstract
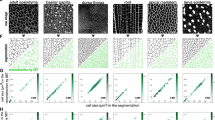
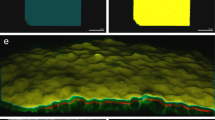
Laboratory automation now commonly allows high-throughput sample preparation, culturing, and acquisition of microscopy images, but quantitative image analysis is often still a painstaking and subjective process. This is a problem especially significant for work on programmed morphogenesis, where the spatial organization of cells and cell types is of paramount importance. To address the challenges of quantitative analysis for such experiments, we have developed TASBE Image Analytics, a software pipeline for automatically segmenting collections of cells using the fluorescence channels of microscopy images. With TASBE Image Analytics, collections of cells can be grouped into spatially disjoint segments, the movement or development of these segments tracked over time, and rich statistical data output in a standardized format for analysis. Processing is readily configurable, rapid, and produces results that closely match hand annotation by humans for all but the smallest and dimmest segments. TASBE Image Analytics can thus provide the analysis necessary to complete the design-build-test-learn cycle for high-throughput experiments in programmed morphogenesis, as validated by our application of this pipeline to process experiments on shape formation with engineered CHO and HEK293 cells.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Lamprecht MR, Sabatini DM, Carpenter AE (2007) Cellprofiler: free, versatile software for automated biological image analysis. BioTechniques 42(1):71–75
Rasband W (2012) Imagej: image processing and analysis in java. Astrophys Sour Code Lib 1:06013
Schindelin J, Rueden CT, Hiner MC, Eliceiri KW (2015) The imagej ecosystem: an open platform for biomedical image analysis. Mol Reprod Dev 82(7-8):518–529
Bajcsy P, Chalfoun J, Simon MH (2018) Web microanalysis of big image data. Springer International Publishing, New York
Stylianidou S, Brennan C, Nissen SB, Kuwada NJ, Wiggins PA (2016) Supersegger: robust image segmentation, analysis and lineage tracking of bacterial cells. Mol Microbiol 102(4):690–700
Clarke ML, Burton RL, Hill AN, Litorja M, Nahm MH, Hwang J (2010) Low-cost, high-throughput, automated counting of bacterial colonies. Cytom Part A 77(8):790–797
Chalfoun J, Majurski M, Dima A, Stuelten C, Peskin A, Brady M (2014) Fogbank: a single cell segmentation across multiple cell lines and image modalities. BMC Bioinformatics 15(1):431
Beal J, Weiss R, Densmore D, Adler A, Appleton E, Babb J, Bhatia S, Davidsohn N, Haddock T, Loyall J, Schantz R, Vasilev V, Yaman F (2012) An end-to-end workflow for engineering of biological networks from high-level specifications. ACS Synth Biol 1(8):317–331
Beal J, Overney C, Adler A, Yaman F, Tiberio L, Samineni M (2019) Tasbe flow analytics: a package for calibrated flow cytometry analysis. ACS Synth Biol 8(7):1524–1529
Efford N (2000) Digital image processing: a practical introduction using java (with CD-ROM), 1st edn. Addison-Wesley Longman Publishing Co., Inc., Boston, MA, USA
Forsyth DA, Ponce J (2002) Computer vision: a modern approach. Professional Technical Reference, Prentice Hall
Tinevez JY, Perry N, Schindelin J, Hoopes GM, Reynolds GD, Laplantine E, Bednarek SY, Shorte SL, Eliceiri KW (2017) Trackmate: an open and extensible platform for single-particle tracking. Methods 115:80–90
Rother C, Kolmogorov V, Blake A (2004) Grabcut: interactive foreground extraction using iterated graph cuts. ACM Trans Graph 23:309–314
Bradski G (2000) The openCV library. Dr Dobb’s J Softw Tools 120:122–125
Everingham M, Van Gool L, Williams CKI, Winn J, Zisserman A (2010) The pascal visual object classes (voc) challenge. Int J Comput Vis 88(2):303–338
Tan PN, Steinbach M, Kumar V (2005) Introduction to data mining, 1st edn. Addison-Wesley Longman Publishing Co., Inc., Boston, MA, USA
Tordoff J, Krajnc M, Walczak N, Lima M, Beal J, Shvartsman S, Weiss R (2020) Incomplete cell sorting creates engineerable structures with long term stability. Cell Reports Physical Science
Acknowledgments
This work has been supported by the Defense Advanced Research Projects Agency under Contract No. W911NF-17-2-0098. The views, opinions, and/or findings expressed are of the author(s) and should not be interpreted as representing official views or policies of the Department of Defense or the U.S. Government. This document does not contain technology or technical data controlled under either U.S. International Traffic in Arms Regulation or U.S. Export Administration Regulations.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Editor(s) (if applicable) and The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Walczak, N., Beal, J., Tordoff, J., Weiss, R. (2021). TASBE Image Analytics: A Processing Pipeline for Quantifying Cell Organization from Fluorescent Microscopy. In: Ebrahimkhani, M.R., Hislop, J. (eds) Programmed Morphogenesis. Methods in Molecular Biology, vol 2258. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1174-6_1
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1174-6_1
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1173-9
Online ISBN: 978-1-0716-1174-6
eBook Packages: Springer Protocols