Abstract
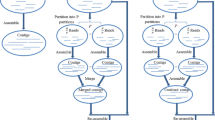
Acquisition of high-quality bacterial genomes is fundamental, while having in mind investigation of subtitle intraspecies variation in addition to development of sensitive species-specific tools for detection and identification of the pathogens. In this view, Pacific Biosciences technology seems highly tempting taking into consideration over 10,000 bp length of the generated reads. In this work, we describe a bacterial genome assembly pipeline based on open-source software that might be handled also by non-bioinformaticians interested in transformation of sequencing data into reliable biological information. With the use of this method, we successfully closed six Dickeya solani genomes, while the assembly process was run just on a slightly improved desktop computer.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Quail M, Smith ME, Coupland P et al (2012) A tale of three next generation sequencing platforms: comparison of Ion Torrent, Pacific Biosciences and Illumina MiSeq sequencers. BMC Genomics 13:341. https://doi.org/10.1186/1471-2164-13-341
Eid J, Fehr A, Gray J et al (2009) Real-time DNA sequencing from single polymerase molecules. Science 323:133–138. https://doi.org/10.1126/science.1162986
Korlach J, Bjornson KP, Chaudhuri BP et al (2010) Real-time DNA sequencing from single polymerase molecules. Methods Enzymol 472:431–455. https://doi.org/10.1016/S0076-6879(10)72001-2
Liolios K, Chen I-MA, Mavromatis K et al (2010) The Genomes On Line Database (GOLD) in 2009: status of genomic and metagenomic projects and their associated metadata. Nucleic Acids Res 38:D346–D354. https://doi.org/10.1093/nar/gkp848
Galardini M, Biondi EG, Bazzicalupo M, Mengoni A (2011) CONTIGuator: a bacterial genomes finishing tool for structural insights on draft genomes. Source Code Biol Med 6:11. https://doi.org/10.1186/1751-0473-6-11
Tettelin H, Riley D, Cattuto C, Medini D (2008) Comparative genomics: the bacterial pan-genome. Curr Opin Microbiol 11:472–477. https://doi.org/10.1016/J.MIB.2008.09.006
Adeolu M, Alnajar S, Naushad S, Gupta RS (2016) Genome-based phylogeny and taxonomy of the ‘Enterobacteriales’: proposal for Enterobacterales ord. nov. divided into the families Enterobacteriaceae, Erwiniaceae fam. nov., Pectobacteriaceae fam. nov., Yersiniaceae fam. nov., Hafniaceae fam. nov., Morganellaceae fam. nov., and Budviciaceae fam. nov. Int J Syst Evol Microbiol 66:5575–5599. https://doi.org/10.1099/ijsem.0.001485
Perombelon MCM, Kelman A (1980) Ecology of the soft rot Erwinias. Annu Rev Phytopathol 18:361–387. https://doi.org/10.1146/annurev.py.18.090180.002045
Toth IK, van der Wolf JM, Saddler G, Lojkowska E, Hellas V, Pirhonen M, Tsror L, Elphinstone JG (2011) Dickeya species: an emerging problem for potato production in Europe. Plant Pathol 60:385–399. https://doi.org/10.1111/j.1365-3059.2011.02427.x
Potrykus M, Golanowska M, Hugouvieux-Cotte-Pattat N, Lojkowska E (2014) Regulators involved in Dickeya solani virulence, genetic conservation, and functional variability. Mol Plant-Microbe Interact 27:700–711. https://doi.org/10.1094/MPMI-09-13-0270-R
Potrykus M, Golanowska M, Sledz W, Zoledowska S, Motyka A, Kołodziejska A, Butrymowicz J, Lojkowska E (2016) Biodiversity of Dickeya spp. isolated from potato plants and water sources in temperate climate. Plant Dis 100:408–417. https://doi.org/10.1094/PDIS-04-15-0439-RE
Zoledowska S, Motyka A, Zukowska D, Sledz W, Lojkowska E (2018) Population structure and biodiversity of Pectobacterium parmentieri isolated from potato fields in temperate climate. Plant Dis 102:154–164. https://doi.org/10.1094/PDIS-05-17-0761-RE
Zoledowska S, Motyka-Pomagruk A, Sledz W, Lojkowska E (2018) High genomic variability in the plant pathogenic bacterium Pectobacterium parmenieri deciphered from de novo assembled complete genomes. BMC Genomics 19:751. https://doi.org/10.1186/s12864-018-5140-9
Golanowska M, Potrykus M, Motyka-Pomagruk A, Kabza M, Bacci G, Galardini M, Bazzicalupo M, Makalowska I, Smalla K, Mengoni A, Hugouvieux-Cotte-Pattat N, Lojkowska E (2018) Comparison of highly and weakly virulent Dickeya solani strains, with a view on the pangenome and panregulon of this species. Front Microbiol 9:1940. https://doi.org/10.3389/fmicb.2018.01940
Golanowska M, Galardini M, Bazzicalupo M, Hugouvieux-Cotte-Pattat N, Mengoni A, Potrykus M, Slawiak M, Lojkowska E (2015) Draft genome sequence of a highly virulent strain of the plant pathogen Dickeya solani, IFB0099. Genome Announc 3:e00109–e00115. https://doi.org/10.1128/genomeA.00109-15
Bacci G, Bazzicalupo M, Benedetti A, Mengoni A (2014) StreamingTrim 1.0: a Java software for dynamic trimming of 16S rRNA sequence data from metagenetic studies. Mol Ecol Resour 14:426–434. https://doi.org/10.1111/1755-0998.12187
Koren S, Schatz MC, Walenz BP et al (2012) Hybrid error correction and de novo assembly of single-molecule sequencing reads. Nat Biotechnol 30:693–700. https://doi.org/10.1038/nbt.2280
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Slawiak M, Łojkowska E, van der Wolf JM (2009) First report of bacterial soft rot on potato caused by Dickeya sp. (syn. Erwinia chrysanthemi) in Poland. Plant Pathol 58:794–794. https://doi.org/10.1111/j.1365-3059.2009.02028.x
Slawiak M, van Beckhoven JRCM, Speksnijder AGCL et al (2009) Biochemical and genetical analysis reveal a new clade of biovar 3 Dickeya spp. strains isolated from potato in Europe. Eur J Plant Pathol 125:245–261. https://doi.org/10.1007/s10658-009-9479-2
Golanowska M, Kielar J, Lojkowska E (2017) The effect of temperature on the phenotypic features and the maceration ability of Dickeya solani strains isolated in Finland, Israel and Poland. Eur J Plant Pathol 147:803–817. https://doi.org/10.1007/s10658-016-1044-1
Degefu Y, Potrykus M, Golanowska M, Virtanen E, Lojkowska E (2013) A new clade of Dickeya spp. plays a major role in potato blackleg outbreaks in North Finland. Ann Appl Biol 162:231–241. https://doi.org/10.1111/aab.12020
Motyka-Pomagruk A (2019) Genotypic and phenotypic characterization of bacteria from Dickeya solani species and development of novel control methods against phytopathogens. PhD thesis, University of Gdańsk
Koren S, Walenz BP, Berlin K et al (2017) Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res 27:722–736. https://doi.org/10.1101/gr.215087.116
Chin CS, Alexander DH, Marks P et al (2013) Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Methods 10:563–569. https://doi.org/10.1038/nmeth.2474
Pacific Biosciences P/N 000-710-821-13 (2014) Template preparation and sequencing guide. Pacific Biosciences, Menlo Park, CA
Pacific Biosciences 100-338-500-01 (2014) Introduction to SMRTbell™ template preparation. Pacific Biosciences, Menlo Park, CA
Li H, Shahriari AR, Wysoker A (2009) Durbin R; 1000 genome project data processing subgroup. The sequence alignment/MAP format and samtools. Bioinformatics 25:2078–2079
Acknowledgments
All sequencing and comparative genomics research tasks were conducted thanks to founding from National Science Centre in Poland via 2014/14/M/NZ8/00501 granted to E.L. A.M.P. is supported from National Science Centre in Poland via 2016/21/N/NZ1/02783.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Motyka-Pomagruk, A., Zoledowska, S., Kabza, M., Lojkowska, E. (2021). PacBio-Based Protocol for Bacterial Genome Assembly. In: Mengoni, A., Bacci, G., Fondi, M. (eds) Bacterial Pangenomics. Methods in Molecular Biology, vol 2242. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1099-2_1
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1099-2_1
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1098-5
Online ISBN: 978-1-0716-1099-2
eBook Packages: Springer Protocols