Abstract
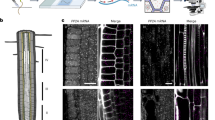
Single-molecule FISH (smFISH) has been widely used in animal tissue to localize and quantify RNAs with high specificity. This protocol describes an smFISH method optimized for highly autofluorescent plant tissue. It provides details on fixation buffers and protocols to protect the integrity of plant samples. We also provide smFISH hybridization conditions to detect plant RNA with ~50 fluorescently labeled DNA oligonucleotides. In addition, this protocol provides instructions on linear spectral unmixing of smFISH signal from background autofluorescence by confocal microscopy and a method to quantify the smFISH spots that reflect the copy number of target RNA.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Anamthawatjonsson K, Reader SM (1995) Pre-annealing of total genomic DNA probes for simultaneous genomic in-situ hybridization. Genome 38(4):814–816
Maluszynska J, Schweizer D (1989) Ribosomal RNA genes in B chromosomes of Crepis capillaris detected by non-radioactive in situ hybridization. Heredity 62(Pt 1):59–65
Fransz PF, Stam M, Montijn B, TenHoopen R, Wiegant J, Kooter JM, Oud O, Nanninga N (1996) Detection of single-copy genes and chromosome rearrangements in Petunia hybrida by fluorescence in situ hybridization. Plant J 9(5):767–774
Weiss H, Pasierbek P, Maluszynska J (2000) An improved nonfluorescent detection system for in situ hybridization in plants. Biotech Histochem 75(2):49–53
Trinh le A, McCutchen MD, Bonner-Fraser M, Fraser SE, Bumm LA, McCauley DW (2007) Fluorescent in situ hybridization employing the conventional NBT/BCIP chromogenic stain. BioTechniques 42(6):756–759. https://doi.org/10.2144/000112476
Taniguchi Y, Choi PJ, Li GW, Chen H, Babu M, Hearn J, Emili A, Xie XS (2010) Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells. Science 329(5991):533–538. https://doi.org/10.1126/science.1188308
Tutucci E, Livingston NM, Singer RH, Wu B (2018) Imaging mRNA in vivo, from birth to death. Annu Rev Biophys 47:85–106. https://doi.org/10.1146/annurev-biophys-070317-033037
Raj A, van den Bogaard P, Rifkin SA, van Oudenaarden A, Tyagi S (2008) Imaging individual mRNA molecules using multiple singly labeled probes. Nat Methods 5(10):877–879
Batish M, Raj A, Tyagi S (2011) Single molecule imaging of RNA in situ. Methods Mol Biol 714:3–13. https://doi.org/10.1007/978-1-61779-005-8_1
Vargas DY, Shah K, Batish M, Levandoski M, Sinha S, Marras SA, Schedl P, Tyagi S (2011) Single-molecule imaging of transcriptionally coupled and uncoupled splicing. Cell 147(5):1054–1065. https://doi.org/10.1016/j.cell.2011.10.024
Rosa S, Duncan S, Dean C (2016) Mutually exclusive sense-antisense transcription at FLC facilitates environmentally induced gene repression. Nat Commun 7:13031
Huang K, Baldrich P, Meyers BC, Caplan JL (2019) sRNA-FISH: versatile fluorescent in situ detection of small RNAs in plants. Plant J 98:359–369. https://doi.org/10.1111/tpj.14210
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B, Tinevez JY, White DJ, Hartenstein V, Eliceiri K, Tomancak P, Cardona A (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9(7):676–682. https://doi.org/10.1038/Nmeth.2019
Mueller F, Senecal A, Tantale K, Marie-Nelly H, Ly N, Collin O, Basyuk E, Bertrand E, Darzacq X, Zimmer C (2013) FISH-quant: automatic counting of transcripts in 3D FISH images. Nat Methods 10(4):277–278
Acknowledgments
This project was supported by the US NSF Plant Genome Research Program, awards 1649424, 1611853, and 1754097. We would like to thank members of the Batish lab for input on single-molecule in situ hybridization, and members of the Meyers and Caplan labs for help and support. Microscopy equipment was acquired with a shared instrumentation grant (S10 OD016361) and access was supported by the NIH-NIGMS (P20 GM103446), the NSF (IIA-1301765), and the State of Delaware.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Huang, K., Batish, M., Teng, C., Harkess, A., Meyers, B.C., Caplan, J.L. (2020). Quantitative Fluorescence In Situ Hybridization Detection of Plant mRNAs with Single-Molecule Resolution. In: Heinlein, M. (eds) RNA Tagging. Methods in Molecular Biology, vol 2166. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0712-1_2
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0712-1_2
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0711-4
Online ISBN: 978-1-0716-0712-1
eBook Packages: Springer Protocols