Abstract
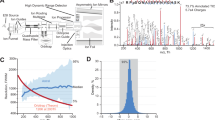
Recent advances in mass spectrometry (MS)-based quantitative proteomics now allow the identification and quantification of deep proteomes and post-translational modifications (PTMs) in relatively short times. Therefore, in the last few years, this technology has proven successful in the circadian field to characterize temporal oscillations of the proteome and more recently PTMs in cellular systems and in tissues. In this chapter, we describe a robust and simple protocol, based on the EasyPhos workflow, to enable preparation of large number of proteomes and phosphoproteomes from mouse tissues for MS-based quantitative analysis. We additionally discuss computational methods to analyze proteome and phosphoproteome time series to determine circadian oscillations.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Panda S et al (2002) Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109(3):307–320
Storch KF et al (2002) Extensive and divergent circadian gene expression in liver and heart. Nature 417(6884):78–83
Zhang R et al (2014) A circadian gene expression atlas in mammals: implications for biology and medicine. Proc Natl Acad Sci U S A 111(45):16219–16224
Masri S et al (2013) Circadian acetylome reveals regulation of mitochondrial metabolic pathways. Proc Natl Acad Sci U S A 110(9):3339–3344
Reddy AB et al (2006) Circadian orchestration of the hepatic proteome. Curr Biol 16(11):1107–1115
Humphrey SJ, Azimifar SB, Mann M (2015) High-throughput phosphoproteomics reveals in vivo insulin signaling dynamics. Nat Biotechnol 33(9):990–995
Sharma K et al (2014) Ultradeep human phosphoproteome reveals a distinct regulatory nature of Tyr and Ser/Thr-based signaling. Cell Rep 8(5):1583–1594
Junger MA, Aebersold R (2014) Mass spectrometry-driven phosphoproteomics: patterning the systems biology mosaic. Wiley Interdiscip Rev Dev Biol 3(1):83–112
Larance M, Lamond AI (2015) Multidimensional proteomics for cell biology. Nat Rev Mol Cell Biol 16(5):269–280
Altelaar AF, Munoz J, Heck AJ (2013) Next-generation proteomics: towards an integrative view of proteome dynamics. Nat Rev Genet 14(1):35–48
Mauvoisin D et al (2014) Circadian clock-dependent and -independent rhythmic proteomes implement distinct diurnal functions in mouse liver. Proc Natl Acad Sci U S A 111(1):167–172
Robles MS, Cox J, Mann M (2014) In-vivo quantitative proteomics reveals a key contribution of post-transcriptional mechanisms to the circadian regulation of liver metabolism. PLoS Genet 10(1):e1004047
Robles MS, Humphrey SJ, Mann M (2017) Phosphorylation is a central mechanism for circadian control of metabolism and physiology. Cell Metab 25(1):118–127
Wang J et al (2017) Nuclear proteomics uncovers diurnal regulatory landscapes in mouse liver. Cell Metab 25(1):102–117
Humphrey SJ, Azimifar SB, Mann M (2015) High-throughput phosphoproteomics reveals in vivo insulin signaling dynamics. Nat Biotechnol 33(9):990–U142
Humphrey SJ, Karayel O, James DE, Mann M (2018) High-throughput and high-sensitivity phosphoproteomics with the EasyPhos platform. Nat Protoc 13(9):1897–1916
Noya SB, Colameo D, Brüning F, Spinnler A, Mircsof D, Opitz L, Mann M, Tyagarajan SK, Robles MS, Brown SA (2019) The forebrain synaptic transcriptome is organized by clocks but its proteome is driven by sleep. Science 366(6462):eaav2642
Brüning F, Noya SB, Bange T, Koutsouli S, Rudolph JD, Tyagarajan SK, Cox J, Mann M, Brown SA, Robles MS (2019) Sleep-wake cycles drive daily dynamics of synaptic phosphorylation. Science 366(6462):eaav3617
Rappsilber J, Ishihama Y, Mann M (2003) Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal Chem 75(3):663–670
Wisniewski JR (2013) Proteomic sample preparation from formalin fixed and paraffin embedded tissue. J Vis Exp (79)
Tyanova S et al (2016) The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat Methods 13(9):731–740
Acknowledgments
We thank Matthias Mann and group members for their support.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Brüning, F., Humphrey, S.J., Robles, M.S. (2021). Phosphoproteome and Proteome Sample Preparation from Mouse Tissues for Circadian Analysis. In: Brown, S.A. (eds) Circadian Clocks. Methods in Molecular Biology, vol 2130. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0381-9_14
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0381-9_14
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0380-2
Online ISBN: 978-1-0716-0381-9
eBook Packages: Springer Protocols