Abstract
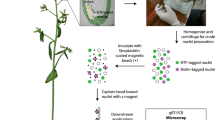
Cells differentiate from undifferentiated precursors in order to establish the tissues of vascular plants. The different cell types and stem cells are first specified in the early embryo. How cell type specification is instructed by transcriptional control on a genome-wide level is poorly understood. A major hurdle has been the technical challenge associated with obtaining cellular transcriptomes in this inaccessible tissue. Recently, we adapted a two-component genetic labeling system called INTACT to isolate nuclei and generate a microarray-based expression atlas of the cell types in the early Arabidopsis thaliana embryo. Here we present a step-by-step description of the adapted INTACT protocol, as well as the approach to generate transcriptomic profiles. This protocol has been adapted to account for using seeds with embryos of various developmental stages as a starting material, and the relatively few cell type-specific nuclei that can be isolated from embryos.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Palovaara J, de Zeeuw T, Weijers D (2016) Tissue and organ initiation in the plant embryo: a first time for everything. Annu Rev Cell Dev Biol 32:47–75
Deal RB, Henikoff S (2010) A simple method for gene expression and chromatin profiling of individual cell types within a tissue. Dev Cell 18:1030–1040
Deal RB, Henikoff S (2011) The INTACT method for cell type–specific gene expression and chromatin profiling in Arabidopsis thaliana. Nat Protoc 6:56–68
Amin NM, Greco TM, Kuchenbrod LM, Rigney MM, Chung M-I, Wallingford JB et al (2014) Proteomic profiling of cardiac tissue by isolation of nuclei tagged in specific cell types (INTACT). Development 141:962–973
Foley SW, Gosai SJ, Wang D, Selamoglu N, Solitti AC, Köster T et al (2017) A global view of RNA-protein interactions reveals novel root hair cell fate regulators. Dev Cell 41:204–220. e5
Henry GL, Davis FP, Picard S, Eddy SR (2012) Cell type–specific genomics of Drosophila neurons. Nucleic Acids Res 40:9691–9704
Mo A, Mukamel EA, Davis FP, Luo C, Henry GL, Picard S et al (2015) Epigenomic signatures of neuronal diversity in the mammalian brain. Neuron 86:1369–1384
Moreno-Romero J, Santos-González J, Hennig L, Köhler C (2017) Applying the INTACT method to purify endosperm nuclei and to generate parental-specific epigenome profiles. Nat Protoc 12:238–254
Park K, Kim MY, Vickers M, Park J-S, Hyun Y, Okamoto T et al (2016) DNA demethylation is initiated in the central cells of Arabidopsis and rice. Proc Natl Acad Sci U S A 113:15138–15143
Reynoso MA, Pauluzzi GC, Kajala K, Cabanlit S, Velasco J, Bazin J et al (2018) Nuclear transcriptomes at high resolution using retooled INTACT. Plant Physiol 176:270–281
Ron M, Kajala K, Pauluzzi G, Wang D, Reynoso MA, Zumstein K et al (2014) Hairy root transformation using agrobacterium rhizogenes as a tool for exploring cell type-specific gene expression and function using tomato as a model. Plant Physiol 166:455–469
Steiner FA, Talbert PB, Kasinathan S, Deal RB, Henikoff S (2012) Cell-type-specific nuclei purification from whole animals for genome-wide expression and chromatin profiling. Genome Res 22:766–777
Palovaara J, Saiga S, Wendrich JR, van ‘t Wout Hofland N, van Schayck JP, Hater F et al (2017) Transcriptome dynamics revealed by a gene expression atlas of the early Arabidopsis embryo. Nat Plants 3:894–904
Belmonte MF, Kirkbride RC, Stone SL, Pelletier JM, Bui AQ, Yeung EC et al (2013) Comprehensive developmental profiles of gene activity in regions and subregions of the Arabidopsis seed. Proc Natl Acad Sci U S A 110:E435–E444
Casson S, Spencer M, Walker K, Lindsey K (2005) Laser capture microdissection for the analysis of gene expression during embryogenesis of Arabidopsis. Plant J 42:111–123
Slane D, Kong J, Berendzen KW, Kilian J, Henschen A, Kolb M et al (2014) Cell type-specific transcriptome analysis in the early Arabidopsis thaliana embryo. Development 141:4831–4840
Lin K, Kools H, de Groot PJ, Gavai AK, Basnet RK, Cheng F et al (2011) MADMAX—management and analysis database for multiple ~omics experiments. J Integr Bioinform 8:160
Wang D, Deal RB (2015) Epigenome profiling of special plant cell types using a streamlined INTACT protocol and ChIP-seq. Methods Mol Biol 1284:3–25
Acknowledgments
The authors thank Tatyana Radoeva and Thomas Nakel for providing the Arabidopsis seed image in Fig. 1a and for image editing in Fig. 1b, respectively. This work was supported by the Federation of European Biochemical Societies (FEBS) to J.P. and by the European Research Council (ERC; Starting Grant “CELLPATTERN”; Contract number 281573) and ERA-CAPS (EURO-PEC; 849.13.006) to D.W.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Palovaara, J., Weijers, D. (2020). Cell Type-Specific Transcriptomics in the Plant Embryo Using an Adapted INTACT Protocol. In: Bayer, M. (eds) Plant Embryogenesis. Methods in Molecular Biology, vol 2122. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0342-0_11
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0342-0_11
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0341-3
Online ISBN: 978-1-0716-0342-0
eBook Packages: Springer Protocols