Abstract
Many RNA architectures were discovered to be involved in essential biological pathways acting as catalysts and/or regulators of gene expression, transcription, translation, splicing, or viral infection. The key to understand their diverse biological functions is to investigate their structure and dynamic. Nuclear Magnetic Resonance (NMR) is a powerful method to gain insight into these properties. However, the study of high-molecular-weight RNAs by NMR remains challenging. Advances in biochemical and NMR methods over the recent years allow to overcome the limitation of NMR. In particular, the incorporation of paramagnetic probes, coupled to the measurement of the induced effects on nuclear spins, has become an efficient tool providing long-range distance restraints and information on dynamic in solution. At the same time, the use of spin label enabled the application of Electron Paramagnetic Resonance (EPR) to study biological macromolecules. Combining NMR and EPR is emerging as a new approach to investigate the architecture of biological systems.
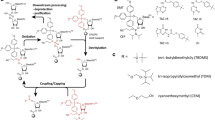
Here, we describe an efficient protocol to introduce a paramagnetic probe into a RNA at a specific position. This method enables various combinations of isotopic labeling for NMR and is also of interest for EPR studies.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Joyce GF (2002) The antiquity of RNA-based evolution. Nature 418:214–221
Kruger K, Grabowski PJ, Zaug AJ, Sands J, Gottschling DE, Cech TR (1982) Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell 31:147–157
Doudna JA, Cech TR (2002) The chemical repertoire of natural ribozymes. Nature 418:222–228
Walter NG (2007) Ribozyme catalysis revisited: is water involved? Mol Cell 28:923–929
Serganov A, Patel DJ (2007) Ribozymes, riboswitches and beyond: regulation of gene expression without proteins. Nat Rev Genet 8:776–790
Mandal M, Breaker RR (2004) Gene regulation by riboswitches. Nat Rev Mol Cell Biol 5:451–463
Grosshans H, Filipowicz W (2008) Molecular biology: the expanding world of small RNAs. Nature 451:414–416
Hannon GJ (2002) RNA interference. Nature 418:244–251
Bevilacqua PC, Blose JM (2008) Structures, kinetics, thermodynamics, and biological functions of RNA hairpins. Annu Rev Phys Chem 59:79–103
Brierley I, Digard P, Inglis SC (1989) Characterization of an efficient coronavirus ribosomal frameshifting signal: requirement for an RNA pseudoknot. Cell 57:537–547
Brunel C, Marquet R, Romby P, Ehresmann C (2002) RNA loop-loop interactions as dynamic functional motifs. Biochimie 84:925–944
Derrigo M, Cestelli A, Savettieri G, Di Liegro I (2000) RNA-protein interactions in the control of stability and localization of messenger RNA (review). Int J Mol Med 5:111–123
Kolb FA, Engdahl HM, Slagter-Jäger JG, Ehresmann B, Ehresmann C, Westhof E, Wagner EG, Romby P (2000) Progression of a loop-loop complex to a four-way junction is crucial for the activity of a regulatory antisense RNA. EMBO J 19:5905–5915
Tomizawa J, Som T (1984) Control of ColE1 plasmid replication: enhancement of binding of RNA I to the primer transcript by the Rom protein. Cell 38:871–878
Allen M, Varani L, Varani G (2001) Nuclear magnetic resonance methods to study structure and dynamics of RNA-protein complexes. Methods Enzymol 339:357–376
Latham MP, Brown DJ, McCallum SA, Pardi A (2005) NMR methods for studying the structure and dynamics of RNA. Chembiochem 6:1492–1505
Fürtig B, Buck J, Manoharan V, Bermel W, Jäschke A, Wenter P, Pitsch S, Schwalbe H (2007) Time-resolved NMR studies of RNA folding. Biopolymers 86:360–383
Getz M, Sun X, Casiano-Negroni A, Zhang Q, Al-Hashimi HM (2007) NMR studies of RNA dynamics and structural plasticity using NMR residual dipolar couplings. Biopolymers 86:384–402
Dominguez C, Schubert M, Duss O, Ravindranathan S, Allain FH-T (2011) Structure determination and dynamics of protein–RNA complexes by NMR spectroscopy. Prog Nucl Magn Reson Spectrosc 58:1–61
Duss O, Lukavsky PJ, Allain FH-T (2012) Isotope labeling and segmental labeling of larger RNAs for NMR structural studies. In: Atreya HS (ed) Isotope labeling in biomolecular NMR. Springer, Dordrecht, pp 121–144
Yadav DK, Lukavsky PJ (2016) NMR solution structure determination of large RNA-protein complexes. Prog Nucl Magn Reson Spectrosc 97:57–81
Schlundt A, Tants J-N, Sattler M (2017) Integrated structural biology to unravel molecular mechanisms of protein-RNA recognition. Methods 118–119:119–136
Barnwal RP, Yang F, Varani G (2017) Applications of NMR to structure determination of RNAs large and small. Arch Biochem Biophys 628:42–56
Cai S, Zhu L, Zhang Z, Chen Y (2007) Determination of the three-dimensional structure of the Mrf2−DNA complex using paramagnetic spin labeling. Biochemistry 46:4943–4950
Hennig J, Warner LR, Simon B, Geerlof A, Mackereth CD, Sattler M (2015) Structural analysis of protein–RNA complexes in solution using NMR paramagnetic relaxation enhancements. Methods Enzymol 558:333–362
Ramos A, Varani G (1998) A new method to detect long-range protein−RNA contacts: NMR detection of electron−proton relaxation induced by nitroxide spin-labeled RNA. J Am Chem Soc 120:10992–10993
Wunderlich CH, Huber RG, Spitzer R, Liedl KR, Kloiber K, Kreutz C (2013) A novel paramagnetic relaxation enhancement tag for nucleic acids: a tool to study structure and dynamics of RNA. ACS Chem Biol 8:2697–2706
Iwahara J, Clore GM (2006) Detecting transient intermediates in macromolecular binding by paramagnetic NMR. Nature 440:1227–1230
Helmling C, Bessi I, Wacker A, Schnorr KA, Jonker HRA, Richter C, Wagner D, Kreibich M, Schwalbe H (2014) Noncovalent spin labeling of riboswitch RNAs to obtain long-range structural NMR restraints. ACS Chem Biol 9:1330–1339
Shelke SA, Sandholt GB, Sigurdsson ST (2014) Nitroxide-labeled pyrimidines for non-covalent spin-labeling of abasic sites in DNA and RNA duplexes. Org Biomol Chem 12:7366–7374
Saha S, Hetzke T, Prisner TF, Sigurdsson ST (2018) Noncovalent spin-labeling of RNA: the aptamer approach. Chem Commun (Camb) 54:11749–11752
Macosko JC, Pio MS, Tinoco I, Shin YK (1999) A novel 5 displacement spin-labeling technique for electron paramagnetic resonance spectroscopy of RNA. RNA 5:1158–1166
Grant GPG, Qin PZ (2007) A facile method for attaching nitroxide spin labels at the 5′ terminus of nucleic acids. Nucleic Acids Res 35:e77
Höbartner C, Sicoli G, Wachowius F, Gophane DB, Sigurdsson ST (2012) Synthesis and characterization of RNA containing a rigid and nonperturbing cytidine-derived spin label. J Org Chem 77:7749–7754
Gophane DB, Endeward B, Prisner TF, Sigurdsson ST (2018) A semi-rigid isoindoline-derived nitroxide spin label for RNA. Org Biomol Chem 16:816–824
Shelke SA, Sigurdsson ST (2012) Site-directed spin labelling of nucleic acids. Eur J Org Chem 2012:2291–2301
Büttner L, Seikowski J, Wawrzyniak K, Ochmann A, Höbartner C (2013) Synthesis of spin-labeled riboswitch RNAs using convertible nucleosides and DNA-catalyzed RNA ligation. Bioorg Med Chem 21:6171–6180
Wawrzyniak-Turek K, Höbartner C (2014) Deoxyribozyme-mediated ligation for incorporating EPR spin labels and reporter groups into RNA. Methods Enzymol 549:85–104
Lebars I, Vileno B, Bourbigot S, Turek P, Wolff P, Kieffer B (2014) A fully enzymatic method for site-directed spin labeling of long RNA. Nucleic Acids Res 42:e117
Duss O, Yulikov M, Jeschke G, Allain FH-T (2014) EPR-aided approach for solution structure determination of large RNAs or protein–RNA complexes. Nat Commun 5:3669
Domnick C, Hagelueken G, Eggert F, Schiemann O, Kath-Schorr S (2019) Posttranscriptional spin labeling of RNA by tetrazine-based cycloaddition. Org Biomol Chem 17:1805.
Qin PZ, Dieckmann T (2004) Application of NMR and EPR methods to the study of RNA. Curr Opin Struct Biol 14:350–359
Duss O, Yulikov M, Allain FHT, Jeschke G (2015) Combining NMR and EPR to determine structures of large RNAs and protein-RNA complexes in solution. Methods Enzymol 558:279–331
Nielsen H (2011) Working with RNA. In: Nielsen H (ed) RNA. Humana Press, Totowa, pp 15–28
Milligan JF, Groebe DR, Witherell GW, Uhlenbeck OC (1987) Oligoribonucleotide synthesis using T7 RNA polymerase and synthetic DNA templates. Nucleic Acids Res 15:8783–8798
Wyatt JR, Chastain M, Puglisi JD (1991) Synthesis and purification of large amounts of RNA oligonucleotides. BioTechniques 11:764–769
Nelissen FHT, van Gammeren AJ, Tessari M, Girard FC, Heus HA, Wijmenga SS (2008) Multiple segmental and selective isotope labeling of large RNA for NMR structural studies. Nucleic Acids Res 36:e89
Romaniuk PJ, Uhlenbeck OC (1983) Joining of RNA molecules with RNA ligase. Methods Enzymol 100:52–59
Bullard DR, Bowater RP (2006) Direct comparison of nick-joining activity of the nucleic acid ligases from bacteriophage T4. Biochem J 398:135–144
Milligan JF, Uhlenbeck OC (1989) Synthesis of small RNAs using T7 RNA polymerase. Methods Enzymol 180:51–62
Scott LG, Hennig M (2008) RNA structure determination by NMR. In: Keith JM (ed) Bioinformatics. Humana Press, Totowa, pp 29–61
Fürtig B, Richter C, Wöhnert J, Schwalbe H (2003) NMR spectroscopy of RNA. Chembiochem 4:936–962
Sambrook J, Fritsch E, Maniatis T (1989) Molecular cloning: a laboratory manual, vol vol. 2, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Fajer PG (2006) Electron spin resonance spectroscopy labeling in peptide and protein analysis. In: Encyclopedia of analytical chemistry. https://doi.org/10.1002/9780470027318.a1609
Oppenheim SF, Buettner GR, Rodgers VGJ (1996) Relationship of rotational correlation time from EPR spectroscopy and protein-membrane interaction. J Membr Sci 118:133–139
Stoll S, Schweiger A (2007) EasySpin: simulating cw ESR spectra. Magn Reson 27:299–321
Stoll S, Schweiger A (2006) EasySpin, a comprehensive software package for spectral simulation and analysis in EPR. J Magn Reson 178:42–55
Etienne E, Le Breton N, Martinho M, Mileo E, Belle V (2017) SimLabel: a graphical user interface to simulate continuous wave EPR spectra from site-directed spin labeling experiments. Magn Reson Chem 55:714–719
Qin PZ, Butcher SE, Feigon J, Hubbell WL (2001) Quantitative analysis of the isolated GAAA tetraloop/receptor interaction in solution: a site-directed spin labeling study. Biochemistry 40:6929–6936
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Vileno, B., Lebars, I. (2020). Site-Specific Spin Labeling of RNA for NMR and EPR Structural Studies. In: Arluison, V., Wien, F. (eds) RNA Spectroscopy. Methods in Molecular Biology, vol 2113. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0278-2_15
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0278-2_15
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0277-5
Online ISBN: 978-1-0716-0278-2
eBook Packages: Springer Protocols