Abstract
Medium-chain esters such as isobutyl acetate (IBAc) and isoamyl acetate (IAAc) are high-volume solvents, flavors, and fragrances. Compared to long-chain esters, these short-chain esters are more volatile and are flavor components of many fruits. For example, IAAc has banana flavor and is widely used as food or beverage additives. Currently, they are mainly produced from petroleum feedstocks. Alternatively, metabolic engineering enables the total biosynthesis of IBAc and IAAc directly from glucose in Escherichia coli. The pathways harnessed the power of natural amino acid biosynthesis. In particular, the native valine and leucine pathways in E. coli were utilized to supply precursors. The key enzyme alcohol O-acyltransferases (AAT) will then catalyze esterification reactions to produce IBAc and IAAc. In vitro biochemical characterization of AAT can provide rational guidance for future enzyme engineering or identify new enzymes for other target substrates. The below protocol provides the detailed description of expression and purification of AAT, in vitro enzymatic assays, direct biosyntheses of IBAc or IAAc in E. coli from glucose, and scale-up production of these valuable products in a benchtop bioreactor.
Access provided by CONRICYT – Journals CONACYT. Download protocol PDF
Similar content being viewed by others
Keywords:
- Alcohol acyltransferase (AAT)
- Biofuel
- Esters
- Metabolic engineering
- Pathway manipulation
- Renewable chemicals
1 Introduction
Petroleum is the material basis of modern society that not only fuels our vehicles but also provides precursors for innumerable consumer products. However, concerns such as growing demands, dwindling reserves, and environmental impacts drive the transition from this petroleum-based economy to a green, bio-based economy. Metabolic engineering is a powerful tool that can facilitate the transition [1–3].
Here, medium-chain esters (C6–C10) are selected as the target compounds for biosynthesis in Escherichia coli [4]. These volatile chemicals have versatile applications. They can be added in foods and beverages as flavor enhancers. For cosmetic and fragrance industries, these esters are used to create fruity or floral aromas. The global market of flavors and fragrances was $25.3 billion in 2014 (http://www.bccresearch.com/). Moreover, esters can be used for coatings, solvents, and potential advanced biofuels. They have high energy content and are fully compatible with existing infrastructure [5]. The industrial production of these esters is currently dominated by Fischer esterification. Fischer esterification is a condensation reaction of a carboxylic acid and an alcohol catalyzed by a strong acid such as sulfuric acid with a reaction temperature between 60 and 110°C. This chemical synthesis process is environmentally unfriendly due to the requirement of petroleum-derived feedstocks, corrosive acid, and high reaction temperature.
Several other processes have been developed, but they all have different limitations. For example, esters extracted from plant materials are often in short supply [6]. Enzymatic synthesis which uses lipase or esterase can catalyze the same condensation reaction as Fischer esterification to produce medium-chain esters. However, the reaction thermodynamically favors hydrolysis of the ester in aqueous solution. Therefore, the production reactions are usually performed in organic solvents such as n-hexane to prevent ester hydrolysis [7, 8]. This makes the process environmentally unfriendly. Alternatively, whole-cell biocatalysis is an attractive approach since the reactions are catalyzed under ambient temperature and in aqueous solution. Recently, there are several studies on the production of isoamyl acetate by supplementing 3-methyl-1-butanol (3 MB) to engineered E. coli overexpressing alcohol O-acyltransferase (AAT) [9–11]. Both pathway and cofactor manipulations were implemented to enhance the production. Similarly, lipase has also been applied by whole-cell biocatalysis to produce isoamyl acetate with supplemented 3 MB [12]. Since 3 MB (a petroleum-derived chemical) was fed to fermentation culture in both cases, these processes are not renewable.
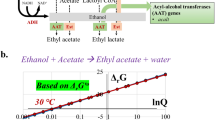
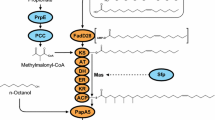
To improve the whole-cell biocatalysis process and make it greener and renewable, here we describe the design and engineering of metabolic pathways for the direct biosynthesis of isobutyl acetate (IBAc) and isoamyl acetate (IAAc) from glucose in E. coli. These pathways exploit amino acid pathways to generate 2-keto acids. The metabolic pathways to IBAc and IAAc production were constructed by expanding native valine and leucine biosynthetic pathways in E. coli as shown in (Fig. 1) (see Note 1 ). The key enzyme to realize the complete pathways is alcohol O-acyltransferase (AAT) which catalyzes the condensation of an alcohol with an acetyl-CoA to form esters. The leaving CoA group makes the reaction step thermodynamically favorable, unlike lipase-catalyzed reactions [13].
Metabolic pathways of the target esters, isobutyl acetate (IBAc) and isoamyl acetate (IAAc) from glucose. Module 1 is valine biosynthesis pathway that converts pyruvate into 2-ketoisovalerate with three enzymes AlsS, IlvC, and IlvD. Module 2 is leucine biosynthesis pathway that adds one more carbon to 2-ketoisovalerate and converts it into 2-keto-4-methyl-pentanoate by LeuABCD. Module 3 represents the last two steps in the Ehrlich pathway which convert the 2-keto acids into their corresponding alcohol. Alcohol O-acyltransferase (AAT) then catalyzes the condensation of an alcohol and an acetyl-CoA to form IBAc or IAAc
Based on the metabolic pathways, four plasmids can be constructed as shown in (Fig. 1). Pathway enzymes that fulfill the direct production of IBAc from glucose are expressed in plasmids pIBAc and pZE-kivd-yqhD (Fig. 2a, c) (see Note 2 ). Pathway enzymes for IAAc production can be expressed in plasmids pIAAc and pZE-kivd-yqhD (Fig. 2b, c). Plasmid pZE-His-AAT (Fig. 2d) (see Note 3 ) is used to produce AAT that can then be purified and used for in vitro enzymatic assays. In this chapter, we describe the expression and purification of AAT, in vitro assays of AAT to determine its kinetic parameters, in vivo production of IBAc and IAAc in shake flasks, and scale-up production of these esters in a 1.3-L benchtop bioreactor.
2 Materials (see Note 4 )
2.1 Plasmid Construction
-
1.
Vectors: pIBAc or pIAAc previously digested with BlpI restriction enzyme. pZE-His-AAT previously digested with BamHI and XbaI restriction enzymes.
-
2.
Primers: BlpI-AAT-For (5′-cgaaagctctctaaGCTGAGCaggagaaattaact-AAT sequence-3′) and BlpI-AAT-Rev (5′-agcctttcgttttatttgatgcctctagaGCTCAGC-AAT sequence-3′) for amplification of AAT. His-AAT-For (5′-gagaggatcgCATCACCATCACCATCAC GGATCC-AAT-3′) and His-AAT-Rev (5′-gactgagcctttcgttttatttgatgccTCTAGA-AAT-3′) for amplification of His-AAT.
-
3.
PCR templates: E. coli K-12 genomic DNA and genomic DNA containing AAT fragment of interest or synthetic AAT fragment codon-optimized for expression in E. coli.
-
4.
Phusion® High-Fidelity PCR Kit (New England Biolabs (http://www.neb.com)).
-
5.
Gibson Assembly® Mater Mix (New England Biolabs (http://www.neb.com)).
-
6.
Quick Ligation™ Kit (New England Biolabs (http://www.neb.com)).
-
7.
Restriction enzymes: FastDigest BlpI, BamHI, and XbaI (Thermo Scientific (www.thermoscientific.com/)).
-
8.
Zymoclean™ Gel DNA Recovery Kit (Zymo Research (https://www.zymoresearch.com/)).
-
9.
Zyppy™ Plasmid Miniprep Kit (Zymo Research (https://www.zymoresearch.com/)).
-
10.
Mix & Go E. coli Transformation Buffer Set (Zymo Research (https://www.zymoresearch.com/)) is used for making chemically competent cells.
-
11.
Bacterial strains: E. coli XL10-Gold strain (Stratagene (http://www.agilent.com/)) is used for cloning.
-
12.
Antibiotics: ampicillin, kanamycin antibiotics solution (see Subheading 2.8).
-
13.
Growth media: super optimal broth (SOB), yeast extract tryptone × 2 (2 × YT) (see Subheading 2.9).
2.2 Protein Expression
-
1.
Vectors: pZE-His-AAT.
-
2.
Bacteria strain: E. coli Rosetta 2(DE3) (EMD Millipore (http://www.merckmillipore.com/)).
-
3.
Antibiotics: ampicillin antibiotic (see Subheading 2.8).
-
4.
Growth media: Luria–Bertani (LB) growth media (see Subheading 2.9).
-
5.
Isopropyl β-d-1-thiogalactopyranoside (IPTG): use 0.5 mM to induce AAT expression (see Subheading 2.8).
2.3 Protein Purification
-
1.
Lysis buffer: 50 mM Tris–HCl, 100 mM NaCl, 10 mM imidazole, 5% glycerol, 1 mM DTT, adjust pH to 7.6.
-
2.
Wash buffer: 50 mM Tris–HCl, 100 mM NaCl, and 25 mM imidazole, adjust pH to 7.6.
-
3.
Elution buffer: 50 mM Tris–HCl, 250 mM NaCl, and 250 mM imidazole, adjust pH to 8.0.
-
4.
Storage buffer: 50 μM Tris–HCl, 2 mM MgSO4, adjust pH to 8.
-
5.
HisPur Ni-NTA resin (Thermo Scientific (www.thermoscientific.com/)). Store at 4°C.
-
6.
Sonication device: Heat Systems Ultrasonics W-225 Sonicator.
-
7.
CrystalCruz™ Chromatography Columns, 1.5 × 15 cm (Santa Cruz Biotechnology, Inc. (http://www.scbt.com/)).
-
8.
Amicon Ultra-15 centrifugal filter devices (Millipore (http://www.emdmillipore.com/)).
-
9.
Quick Start Bradford protein assay kit (Bio-Rad (www.bio-rad.com/)).
-
10.
Glycerol (Sigma (http://www.sigmaaldrich.com)): prepare 87 v% solution; sterilize by autoclave.
2.4 Enzyme Assay
-
1.
Assay buffer: 100 mM KCl, 2 mM MgCl2, 50 mM Tris–HCl in water, adjust pH to 8.0. (All the reagents were purchased from Sigma (http://www.sigmaaldrich.com).)
-
2.
Acetyl coenzyme A sodium salt solution (Sigma (http://www.sigmaaldrich.com)): 10 mM in water. Store at −80°C.
-
3.
Isobutanol (BioUltra, for molecular biology, Sigma (http://www.sigmaaldrich.com)).
-
4.
3-Methyl-1-butanol (anhydrous, Sigma (http://www.sigmaaldrich.com)).
-
5.
5,5′-Dithiobis-(2-nitrobenzoic acid) (DTNB) solution (Sigma (http://www.sigmaaldrich.com)): 1 mM in water, make fresh as required.
-
6.
Greiner UV-Star® 96-well plates (Sigma (http://www.sigmaaldrich.com)).
2.5 Shake Flask Fermentation
-
1.
Bacteria strain: Wild-type (WT) E. coli indicates strain BW25113.
-
2.
5 × M9 salts: 64 g Na2HPO4 · H2O, 15 g KH2PO4, 2.5 g NaCl, 5.0 g NH4Cl/L water. (All the reagents were purchased from Sigma (http://www.sigmaaldrich.com)).
-
3.
Yeast extract solution: 5 g yeast extract/L.
-
4.
Glucose solution: 400 g glucose/L.
-
5.
Antibiotics: ampicillin, kanamycin antibiotics (see Subheading 2.8).
-
6.
Thiamine hydrochloride solution (Sigma (http://www.sigmaaldrich.com)): 10 mg/mL in sterile double-distilled (Milli-Q) water, filtered. Store at −20°C.
-
7.
Modified trace metal solution (see Subheading 2.8).
-
8.
CaCO3 (Sigma (http://www.sigmaaldrich.com)).
-
9.
Parafilm® M (Sigma (http://www.sigmaaldrich.com)).
2.6 Benchtop Bioreactor Fermentation
-
1.
Glucose feeding solution: 600 g glucose, 7.4 g K2HPO4, 1 mL modified trace metal solution/L water, sterilize by autoclave.
-
2.
28 w/v% NH4OH solution (Sigma (http://www.sigmaaldrich.com)).
-
3.
Bioreactor media: 7.5 g K2HPO4, 2 g MgSO4 · 7H2O, 2 g citric acid · H2O, 0.3 g ferric ammonium citrate, 20 g yeast extract, 0.8 mL 98% sulfuric acid, 1 mL modified trace metal solution, 1 mL vitamin solution/L water.
-
4.
Eppendorf BioFlo® 115 benchtop bioreactor and fermentor (Eppendorf (http://www.eppendorf.com/)).
-
5.
Growth media: Luria–Bertani (LB) growth media (see Subheading 2.9).
-
6.
Modified trace metal solution (see Subheading 2.8).
-
7.
Vitamin solution: thiamine 1 g hydrochloride, 1 g d-(+)-biotin, 1 g nicotinic acid, 4 g pyridoxine hydrochloride/L water.
-
8.
Oleyl alcohol (Sigma (http://www.sigmaaldrich.com)).
2.7 Metabolite Detection
-
1.
Mobile phase: 5 mM H2SO4 in water (Sigma (http://www.sigmaaldrich.com)).
-
2.
Hexane (Sigma (http://www.sigmaaldrich.com)) used for extraction.
-
3.
Agilent 1260 Infinity HPLC with a differential refractive detector (RID) (Agilent Technologies (http://www.agilent.com)).
-
4.
HPLC column: Aminex HPX 87H column (Bio-Rad (www.bio-rad.com)).
-
5.
Hewlett Packard (HP) 6890 gas chromatograph (GC) equipped with a flame ionization detector (FID) (Agilent Technologies (http://www.agilent.com)).
-
6.
GC column: DB-WAX capillary column (30 m, 0.32 mm inside diameter, 0.50 μm film thickness) (Agilent Technologies (http://www.agilent.com)).
2.8 General Buffers and Reagents
-
1.
Ampicillin (Fisher Scientific (http://www.fishersci.com/)): 100 mg/mL in water. Store at −20°C.
-
2.
Kanamycin (VWR (http://us.vwr.com/)): 50 mg/mL in water. Store at −20°C.
-
3.
Isopropyl β-d-1-thiogalactopyranoside (IPTG) (Fisher Scientific (http://www.fishersci.com/)): 1 M in sterile double-distilled (Milli-Q) water stored in 0.5 mL aliquots at −20°C.
-
4.
Modified trace metal solution: 10 g NaCl, 40 g citric acid, 1 g ZnSO4 · 7H2O, 30 g MnSO4 · H2O, 30; 0.1 g CuSO4 · 5H2O; 0.1 g H3BO3, 0.1 g Na2MoO4 · 2H2O, 1 g FeSO4 · 7H2O, 1 g CoCl2 · 6H2O/L water. Sterilize by autoclave.
2.9 Bacteria Growth Media
-
1.
2 × YT: 16 g tryptone plus, 10 g yeast extract, 5 g NaCl/L water.
-
2.
LB: 10 g tryptone plus, 5 g yeast extract, 10 g NaCl/L water.
-
3.
SOB: 20 g tryptone plus, 5 g yeast extract, 0.6 g NaCl, 0.186 g KCl, 2.4 g MgSO4/L water, adjust pH to 7; sterilize by autoclave.
To prepare solid media using Petri dishes, agar at the final concentration of 1.5% was added to the LB media. After the autoclave, wait until the media cool down to about 50°C. Then add antibiotics as desired. Final concentrations of the antibiotics used in this study are as follows: ampicillin 100 μg/mL and kanamycin 50 μg/mL.
3 Methods
3.1 Plasmid Construction
-
1.
The AAT of interest should be amplified by PCR using BlpI-AAT-For and BlpI-AAT-Rev primers (see Subheading 2.1). His-AAT is amplified by PCR using primers His-AAT-For and His-AAT-Rev (see Subheading 2.1). The PCR reaction conditions are 98°C for 30 s, 30 cycles of 98°C for 10 s and 72°C for 1 min, and a final extension of 72°C for 5 min. The reaction volume is 50 μL.
-
2.
Purify the resulted PCR product using a gel DNA recovery kit (see Note 5 ).
-
3.
Clone the AAT insert into previously BlpI-digested pIBAc or pIAAc vector by Gibson assembly. Clone the His-AAT insert into previously BamHI- and XbaI-digested pZE-His-AAT vector by Gibson assembly.
-
4.
Transform the assembly product into XL10-Gold competent cells prepared with SOB and Mix & Go E. coli Transformation Buffer Set.
3.2 Protein Expression
-
1.
Transform Rosetta 2 (DE3) competent cells with pZE-HisAAT vectors and plate the transformed cells on LB-agar plates containing 100 mg/L ampicillin.
-
2.
Leave the plates at 37°C overnight (about 12–16 h) until colonies of transformed bacteria are clearly visible.
-
3.
Overnight cultures with colonies streaked off the plates are inoculated 1% in 200 mL 2 × YT containing 100 mg/L ampicillin in a 500 mL baffled Erlenmeyer flask.
-
4.
Grow cells to OD600 = 0.6–1.0 (about 3–4 h) at 37°C in a rotary shaker (250 rpm).
-
5.
Add IPTG to the culture to 0.5 m final concentration to induce protein expression.
-
6.
Continue protein production in a rotary shaker (250 rpm) at 30°C for 4 h.
-
7.
Pellet the cells by centrifugation at 3,220 rcf for 15 min. For centrifugation, we use 5810 R refrigerated centrifuge (Eppendorf (http://www.eppendorf.com/)).
-
8.
Cell pellet can be stored at −80°C for several weeks or you can immediately proceed to the cell lysis in protein purification.
3.3 Protein Purification
-
1.
All the following steps are carried out on ice or at 4°C to prevent protein degradation.
-
2.
Resuspend the cell pellet with 15 mL lysis buffer (see Subheading 2.3).
-
3.
Lyse cells by sonication. Sonicate each sample with six 1 min cycles with intermittent 1 min rest on ice.
-
4.
Centrifuge at 10,733 rcf for 15 min at 4°C to remove the insoluble part of the cell lysate. For centrifugation, we use 5810 R refrigerated centrifuge (Eppendorf (http://www.eppendorf.com/)).
-
5.
Transfer the soluble fraction to a clean 50 mL conical tube and keep it on ice.
-
6.
Prepare the gravity chromatography column by loading 4 mL HisPur Ni-NTA resin slurry.
-
7.
Allow 20% ethanol (used to make the 50% slurry of Ni-NTA resin) to pass through by gravity to get a 2 mL final resin bed volume.
-
8.
Equilibrate the beads with 10 mL lysis buffer. Allow buffer to drain from the column.
-
9.
Add the prepared protein extract to the resin and collect the flow-through in a 15 mL conical tube.
-
10.
Reapply the flow-through to the resin bed once to maximize binding.
-
11.
Wash the column twice with 20 mL of wash buffer.
-
12.
Elute the bound AAT protein with 12 mL elution buffer. Add the eluted protein in an Amicon Ultra centrifugal filter.
-
13.
Centrifuge at 5,000 rcf for 50 min or more at 4°C until the eluted protein is concentrated down to 0.5 mL or lower.
-
14.
Add storage buffer to 15 mL. Centrifuge at 10,733 rcf for 50 min or more at 4°C until the eluted protein is concentrated down to 0.5 mL.
-
15.
Add 87% glycerol to the purified protein and make it final 20% glycerol. Divide the purified protein into 100 μL aliquots into PCR tubes.
-
16.
Flash frozen with dry ice and methanol mixture and store at −80°C.
-
17.
Analyze the purified AAT by Bradford protein assay and SDS-PAGE.
3.4 Enzyme Assay
-
1.
Prepare ten reaction mixtures with assay buffer (see Subheading 2.4), 10 μL of 10 mM acetyl-CoA solution, 10 μL of the purified AAT solution at a concentration of 1 μM, and 0, 0.5, 1, 2, 3, 5, 10, 20, 50, or 100 mM target alcohol (isobutanol or 3-methyl-1-butanol) to a total volume of 100 μL each (see Note 6 ).
-
2.
Place the reaction mixtures at 30°C for 30 min (see Note 7 ).
-
3.
Stop the reaction by adding 0.06 g NaCl solid powder.
-
4.
Add 0.2 mL 1 mM DTNB solution. Mix well and take 100 μL of the mixture and put it into a 96-well plate.
-
5.
Quantify the yellow 5-thio-2-nitrobenzoate anion product by measuring light absorbance at 412 nm.
-
6.
Adjust the values for unspecific hydrolysis by deducting the absorbance of the controlled sample with no addition of alcohols.
-
7.
Apply a molar extinction coefficient of 13,600 M−1 cm−1 to calculate the reaction rate.
-
8.
Use the nlinfit function in Matlab to calculate the K m and k cat values of the AAT.
3.5 Shake Flask Fermentation
-
1.
Transform WT competent cells with pZE-kivd-yqhD and pIBAc vector set (for production of IBAc) or with pZE-kivd-yqhD and pIAAc vector set (for production of IAAc) and plate the transformed cell on an LB plate containing: hundred 100 μg/mL ampicillin and 50 μg/mL kanamycin. Leave at 37°C overnight (12–16 h) until colonies are clearly visible.
-
2.
Streak off three independent colonies from the plate with freshly transformed cells. Inoculate each colony in 2 mL 2 × YT + 100 μg/mL ampicillin and 50 μg/mL kanamycin in a test tube. Grow shaking (250 rpm) at 37°C overnight (12–16 h).
-
3.
Prepare 125 mL screw cap conical flasks for fermentation by adding 0.3 g CaCO3 into each flask and sterilize them with 121°C for 25 min.
-
4.
Transfer 200 μL of the overnight cultures into the sterilized conical flasks containing 2 mL 5 × M9 salts, 0.5 mL glucose solution, 7.5 mL yeast extract solution, 10 μg/mL thiamine, 0.1 mM IPTG, 100 μg/mL ampicillin, and 50 μg/mL kanamycin.
-
5.
Seal the caps with Parafilm. Place the flasks in a 30°C incubator, gently shaking at 250 rpm for 48 h (see Note 8 ).
-
6.
Collect 1 mL samples and spin down the cells by centrifugation at 20,000 rcf for 15 min and take cell supernatant for analyses. For centrifugation we use Eppendorf Centrifuge 5424 (Eppendorf (http://www.eppendorf.com/)).
3.6 Benchtop Bioreactor Fermentation
-
1.
Calibrate pH meter with pH = 7.0 and 10.0 standard buffer. Add 0.5 L bioreactor media into a 1.3 L benchtop bioreactor. Place the caps on the dO2 and pH meters and bearing housing. Sterilize by autoclave (see Note 9 ).
-
2.
After the medium cool down, adjust the temperature to 37°C by heat blanket and pH to 6.8 by 26% NH4OH solution. Then, add 12.5 mL 40 wt% glucose solution to make a starting 10 g/L glucose concentration. Maintain the airflow rate at 0.5 vvm throughout the fermentation process.
-
3.
Connect the exhaust from the bioreactor to three traps for the ester production as follows: the first one contains 1,800 mL of water cooled with ice–salt mixture, and the second trap is the same as the first one, and the third trap contains 500 mL of oleyl alcohol cooled with tap water.
-
4.
Inoculate transformed production strain in 2 mL 2 × YT + 100 μg/mL ampicillin and 50 μg/mL kanamycin in a test tube. Grow shaking (250 rpm) at 37°C overnight (12–16 h).
-
5.
Transfer 0.5 mL of the overnight pre-culture into 50 mL LB medium + 100 μg/mL ampicillin and 50 μg/mL kanamycin in a 250 mL shake flask.
-
6.
Grow shaking (250 rpm) at 37°C to an OD600 = 1.0–1.2. Add the 50 mL inoculums into the bioreactor. Grow the cells at 37°C and 20% dO2 level to OD600 reach 6.
-
7.
Reduce the temperature to 30°C and dO2 level to 10% (by decreasing the agitation rate). Add 150 μL of 1 M IPTG solution to induce protein expression.
-
8.
Maintain the dO2 level at 10% by adjusting agitation rate between 300 and 800 rpm.
-
9.
Run the fermentation in batch mode until the glucose is totally consumed, and then manually feed glucose feeding solution to meet metabolic demands. Collect samples from the bioreactor and the three traps every 6–12 h and analyze the production of the target ester and other byproducts (see Note 10 ).
3.7 Metabolite Detection
-
1.
Analyze the concentrations of glucose and other byproducts using HPLC-RID. Use 5 mM H2SO4 as the mobile phase at a flow rate of 0.6 mL/min for 40 min. Keep the column temperature at 35°C and RID detector temperature at 50°C. The injection volume is 20 μL.
-
2.
Analyze the concentration of the target ester product (IBAc or IAAc) using GC-FID. Use 2.5 mL hexane to extract from 5 mL cell culture. Mix the mixture for 1 min using vortex, then centrifuge the mixture at 20,000 rcf for 5 min. Take 0.5–1 mL hexane extract for GC analyses.
-
3.
Inject 2 μL of the hexane extract at a 15:1 split ratio. Hold GC oven temperature at 70°C for 2 min and increase it to 120°C with a gradient of 30°C/min. Then increase the oven temperature to 140°C with a gradient of 10°C/min and to 200°C with a 30°C/min gradient. Hold at 200°C for 5 min to clean any remaining chemicals in the column. Hold the temperature of the injector and the FID detector at 250°C.
4 Notes
-
1.
Information about the genes of pathway enzymes are as follows: acetolactate synthase (alsS, from Bacillus subtilis) [14], 2,3-dihydroxy isovalerate oxidoreductase (ilvC, from E. coli), 2,3-dihydroxy isovalerate dehydratase (ilvD, from E. coli) [15], 2-keto-acid decarboxylases (kivd, from Lactococcus lactis) [16], alcohol dehydrogenases (yqhD, from E. coli) [17], 2-isopropylmalate synthase (leuA, from E. coli), isopropylmalate isomerase complex (leuC and leuD, from E. coli), and 3-isopropylmalate dehydrogenase (leuB, from E. coli).
-
2.
Based on our previous experience, endogenous activity of IlvC is good enough for the production of isobutanol; therefore, we did not overexpress IlvC in the plasmid. Similarly, acetyl-CoA is ubiquitous and is not the limiting substrate for IBAc and IAAc production. Thus, we did not overexpress any acetyl-CoA production enzyme.
-
3.
The DNA sequence of the 6 × His tag is 5′-catcaccatcaccatcac-3′.
-
4.
Names of vendors are provided in the list of Materials (Subheading 2) from which we currently purchase chemicals. However, it does not mean that we endorse these particular vendors.
-
5.
Unless mentioned otherwise, standard protocols of the commercially obtained kit should be followed during cloning and purification processes.
-
6.
Use a multi-channel pipette to add the acetyl-CoA solution at last to start all the reactions at the same time for a more accurate measurement. We performed the reaction in 1.5 mL centrifuge tubes.
-
7.
The appropriate reaction time to characterize enzyme kinetics depends on the activity of AAT. Therefore, 30 min is a recommended reaction time, but the optimal time needs to be tested by trial experiments.
-
8.
It is important to seal the caps tightly with Parafilm to prevent the volatile products from escaping out of the flasks.
-
9.
Before sterilization, make sure all the caps have been securely put on the desired positions to prevent the steam from ruining the sensitive components. Do not disconnect the dO2 probe for longer than 10 s after autoclave. Do not autoclave glucose along with the bioreactor media, and add sterilized glucose solution separately.
-
10.
The productivities of the target esters can be characterized as g IBAc/(L · g cells · h) and g IAAc/(L · g cells · h).
References
Connor MR, Liao JC (2009) Microbial production of advanced transportation fuels in non-natural hosts. Curr Opin Biotechnol 20:307–315
Keasling JD (2010) Manufacturing molecules through metabolic engineering. Science 330:1355–1358
Stephanopoulos G (2007) Challenges in engineering microbes for biofuels production. Science 315:801–804
Tai YS, Xiong M, Zhang K (2015) Engineered biosynthesis of medium-chain esters in Escherichia coli. Metab Eng 27:20–28
Lange JP, Price R, Ayoub PM et al (2010) Valeric biofuels: a platform of cellulosic transportation fuels. Angew Chem Int Ed 49:4479–4483
Hari Krishna S, Divakar S, Prapulla SG, Karanth NG (2001) Enzymatic synthesis of isoamyl acetate using immobilized lipase from Rhizomucor miehei. J Biotechnol 87:193–201
Eisenmenger M, Reyes-De-Corcuera J (2010) Kinetics of lipase catalyzed isoamyl acetate synthesis at high hydrostatic pressure in hexane. Biotechnol Lett 32:1287–1291
Ozyilmaz G, Yağız E (2012) Isoamylacetate production by entrapped and covalently bound Candida rugosa and porcine pancreatic lipases. Food Chem 135:2326–2332
San KY, Bennett GN, Berrios-Rivera SJ et al (2002) Metabolic engineering through cofactor manipulation and its effects on metabolic flux redistribution in Escherichia coli. Metab Eng 4:182–192
Vadali RV, Bennett GN, San KY (2004) Applicability of CoA/acetyl-CoA manipulation system to enhance isoamyl acetate production in Escherichia coli. Metab Eng 6:294–299
Vadali RV, Horton CE, Rudolph FB, Bennett GN, San KY (2004) Production of isoamyl acetate in ackA-pta and/or ldh mutants of Escherichia coli with overexpression of yeast ATF2. Appl Microbiol Biotechnol 63:698–704
Brault G, Shareck F, Hurtubise Y, Lepine F, Doucet N (2014) Short-chain flavor ester synthesis in organic media by an E coli whole-cell biocatalyst expressing a newly characterized heterologous lipase. PLoS One 9:e91872
Rodriguez GM, Tashiro Y, Atsumi S (2014) Expanding ester biosynthesis in Escherichia coli. Nat Chem Biol 10:259–265
Renna MC, Najimudin N, Winik LR, Zahler SA (1993) Regulation of the Bacillus Subtilis alsS, alsD, and alsR genes involved in post-exponential-phase production of acetoin. J Bacteriol 175:3863–3875
Daniels DL, Plunkett G, Burland V, Blattner FR (1992) Analysis of the Escherichia coli genome DNA sequence of the region from 84.5 to 86.5 minutes. Science 257:771–778
de la Plaza M, de Palencia PF, Pelaez C, Requena T (2004) Biochemical and molecular characterization of alpha-ketoisovalerate decarboxylase, an enzyme involved in the formation of aldehydes from amino acids by Lactococcus lactis. FEMS Microbiol Lett 238:367–374
Sulzenbacher G, Alvarez K, van den Heuvel RHH et al (2004) Crystal structure of E. coli alcohol dehydrogenase YqhD: evidence of a covalently modified NADP coenzyme. J Mol Biol 342:489–502
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer-Verlag Berlin Heidelberg
About this protocol
Cite this protocol
Tai, YS., Zhang, K. (2015). Designing Bacteria to Produce Esters. In: McGenity, T., Timmis, K., Nogales Fernández, B. (eds) Hydrocarbon and Lipid Microbiology Protocols. Springer Protocols Handbooks. Springer, Berlin, Heidelberg. https://doi.org/10.1007/8623_2015_143
Download citation
DOI: https://doi.org/10.1007/8623_2015_143
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-662-49126-3
Online ISBN: 978-3-662-49127-0
eBook Packages: Springer Protocols