Abstract
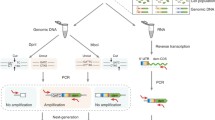
The protocols described help studying the expression of specific genes with the equipment and expertise that is usually available to most research laboratories. They are specifically intended for bacteria grown in pure cultures. The first two protocols are useful to analyse gene expression in vivo and rely on the use fusions to reporter genes such as lacZ and gfp. The third protocol requires purification of total RNA from cells and is based on the transformation of the RNA to a complementary DNA, which is then quantified by a real-time polymerase chain reaction (RT-PCR). It therefore serves to measure the abundance of specific RNAs, or changes in the levels of particular RNAs, under two different conditions. The methods described can answer different questions on the expression of a given gene and therefore complement each other.
The authors Sofía Hernández-Arranz, Ruggero La Rosa, Renata Moreno, Emma Sevilla and Luis Yuste have contributed equally to this work.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Slauch JM, Silhavy TJ (1991) Genetic fusions as experimental tools. Methods Enzymol 204:213–248
Hughes KT, Maloy SR (2007) Use of operon and gene fusions to study gene regulation in Salmonella. Methods Enzymol 421:140–158
Miller WG, Lindow SE (1997) An improved GFP cloning cassette designed for prokaryotic transcriptional fusions. Gene 191:149–153
Stewart GS, Williams P (1992) lux genes and the applications of bacterial bioluminescence. J Gen Microbiol 138:1289–1300
Pannuri A, Yakhnin H, Vakulskas CA, Edwards AN et al (2012) Translational repression of NhaR, a novel pathway for multi-tier regulation of biofilm circuitry by CsrA. J Bacteriol 194:79–89
de Lorenzo V, Herrero M, Jakubzik U, Timmis KN (1990) Mini-Tn5 transposon derivatives for insertion mutagenesis, promoter probing, and chromosomal insertion of cloned DNA in gram-negative eubacteria. J Bacteriol 172:6568–6572
Silva-Rocha R, Martínez-García E, Calles B, Chavarría M et al (2013) The Standard European Vector Architecture (SEVA): a coherent platform for the analysis and deployment of complex prokaryotic phenotypes. Nucleic Acids Res 41:D666–D675
de Lorenzo V, Timmis KN (1994) Analysis and construction of stable phenotypes in gram-negative bacteria with Tn5- and Tn10-derived minitransposons. Methods Enzymol 235:386–405
de Lorenzo V, Fernández JM (2000) Expression vectors and delivery systems. Playing alien genes in remote theaters. Curr Opin Biotechnol 11:427–428
Silva-Rocha R, de Lorenzo V (2011) A composite feed-forward loop I4-FFL involving IHF and Crc stabilizes expression of the XylR regulator of Pseudomonas putida mt-2 from growth phase perturbations. Mol Biosyst 7:2982–2990
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor. NY
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCt method. Methods 25:402–408
Cowles CE, Nichols NN, Harwood CS (2000) BenR, a XylS homologue, regulates three different pathways of aromatic acid degradation in Pseudomonas putida. J Bacteriol 182:6339–6346
Hernández-Arranz S, Moreno R, Rojo F (2013) The translational repressor Crc controls the Pseudomonas putida benzoate and alkane catabolic pathways using a multi-tier regulation strategy. Environ Microbiol 15:227–241
Rojo F (2010) Carbon catabolite repression in Pseudomonas: optimizing metabolic versatility and interactions with the environment. FEMS Microbiol Rev 34:658–684
Acknowledgements
Work was funded by grant BFU2012-32797 from the Spanish Ministry of Economy and Competitiveness.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Berlin Heidelberg
About this protocol
Cite this protocol
Hernández-Arranz, S., La Rosa, R., Moreno, R., Sevilla, E., Yuste, L., Rojo, F. (2014). Protocols on Regulation of Gene Expression. In: McGenity, T., Timmis, K., Nogales , B. (eds) Hydrocarbon and Lipid Microbiology Protocols. Springer Protocols Handbooks. Springer, Berlin, Heidelberg. https://doi.org/10.1007/8623_2014_13
Download citation
DOI: https://doi.org/10.1007/8623_2014_13
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-662-50433-8
Online ISBN: 978-3-662-50435-2
eBook Packages: Springer Protocols