Abstract
Externalizing problems generally refer to a constellation of behaviors and/or disorders characterized by impulsive action and behavioral disinhibition. Phenotypes on the externalizing spectrum include psychiatric disorders, nonclinical behaviors, and personality characteristics (e.g. alcohol use disorders, other illicit substance use, antisocial behaviors, risky sex, sensation seeking, among others). Research using genetic designs including latent designs from twin and family data and more recent designs using genome-wide data reveal that these behaviors and problems are genetically influenced and largely share a common genetic etiology. Large-scale gene-identification efforts have started to identify robust associations between genetic variants and these phenotypes. However, there is still considerable work to be done. This chapter provides an overview of the current state of research into the genetics of behaviors and disorders on the externalizing spectrum.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Externalizing problems refer to a constellation of behaviors and/or disorders characterized by impulsive action and/or behavioral undercontrol. Externalizing problems can be contrasted with internalizing problems in that they typically reflect actions in the external world, rather than internalized processes within the self, such as anxiety, depression, or negative affect. Externalizing problems include a variety of behaviors such as alcohol or substance misuse, antisocial behaviors, aggression, and risk taking (Krueger et al. 2002; Salvatore and Dick 2018; Young et al. 2000).
Problems associated with externalizing behaviors have high social costs. Substance misuse remains one of the leading contributors to preventable mortality and morbidity worldwide. In 2016, alcohol use contributed 4.2% of the total global burden of disease; other drug use contributed to 1.3% of the total global disease burden; and smoking contributed to approximately 12% of all deaths (Degenhardt et al. 2013; Reitsma et al. 2017). In 2017, over 47,000 Americans died as the result of an opioid overdose (Center for Disease Control and Prevention 2019). In addition to the health consequences, these behaviors have significant financial costs. Each year, excessive alcohol use is estimated to cost the United States $250 billion (Sacks et al. 2015). Illicit drugs cost the United States approximately $190 billion (National Drug Intelligence Center 2011) annually, of which $78.5 billion is due to opioid use alone (Florence et al. 2016). And while difficult to calculate, the total cost of crime in the United States is estimated between $690 billion to $3.41 trillion annually (Maurer 2017). Understanding the etiology of these behaviors is of utmost importance for practitioners and policy makers to effectively design prevention and intervention efforts.
2 Epidemiology of Externalizing Behaviors/Disorders
Behaviors and disorders across the externalizing spectrum are highly prevalent. The 12-month prevalence for substance use disorders (SUD) in the United States is approximately 14% for alcohol use disorders (AUD) and 4% for other substance use disorders (SUD), while the lifetime prevalence is much higher (~29% for AUD and ~10% for SUD) (Grant et al. 2015, 2016). These disorders typically manifest during young adulthood, with mean ages of onset ranging from 23.9 for SUD to 26.2 for AUD (Grant et al. 2015, 2016). The lifetime prevalence for other psychiatric disorders related to impulse control, such as attention hyperactivity deficit disorder (ADHD) and conduct disorder (CD), are 5.1% and 9.5%, respectively, and these appear at earlier ages (7 and 13 years old, respectively) (Kessler et al. 2005a; Polanczyk et al. 2007). Taken together, the prevalence for any disorder related to impulse control is high: 24.8%, with a median age of onset = 11 years old (Kessler et al. 2005a). Importantly, substance use and impulse control disorders do not manifest in isolation and show strong comorbidity in past 12-month diagnoses (Grant et al. 2015, 2016; Kessler et al. 2005b). Longitudinal analyses reveal that many externalizing problems, including heavy alcohol use (Chen and Jacobson 2012), illicit drug use (Chen and Jacobson 2012), and antisocial behaviors (Powell et al. 2010), increase across adolescence into young adulthood, followed by a steady decline. Overall, behaviors on the externalizing spectrum are common, with significant variation across the life course.
2.1 Genetic Epidemiology of the Externalizing Spectrum
Twin and family designs use information from close relatives to estimate the heritabilityFootnote 1 of a trait. Twin studies allow researchers to decompose the variance in a trait into additive genetic, shared environmental, and unique environmental influences by comparing the phenotypic correlations of monozygotic (MZ) and dizygotic (DZ) twin pairs. We can estimate these variances due to the fact that MZ twins share all of their genetic variation, while DZ twins share half of their genetic variation, on average. Shared environmental influences, which refer to environments that make twins more similar, include conditions such as neighborhood context, family socioeconomic status, and religion. Unique environmental influences refer to experiences that have the effect of making twins more different from each other than expected based on their genetic sharing, for example, if one twin experiences a trauma, or has a different peer group. When the within-pair MZ correlation for a phenotype is larger than the within-pair DZ correlation, this suggests the importance of genetic influences on the trait under study. When the DZ correlation for a phenotype is more than half of the MZ correlation, this suggests the presence of shared environmental influences. When the MZ correlation is less than unity, unique environmental influences are inferred (measurement error is also confounded with unique environmental influences) (Neale and Cardon 2013).
Many of the individual phenotypes on the externalizing spectrum demonstrate modest to considerable heritability (h 2). SUD have moderate genetic influences, with ~50% of the variance in AUD (Verhulst et al. 2015), 50–60% of the variance in problematic cannabis use (Verweij et al. 2010), ~40–80% of the variance in cocaine use disorders (Kendler et al. 2000, 2003a), 20–50% of the variation in opioid dependence (Kendler et al. 2003a; van den Bree et al. 1998), and ~60% of the variance in nicotine dependence (Maes et al. 2004) being due to genetic influences (h 2). Related psychiatric and behavioral outcomes, such as ADHD (h 2 = 74%), antisocial behavior (h 2 = 32%), rule breaking (h 2 = 48%), and aggression (h 2 = 65%), are moderately-to-strongly heritable (Burt 2009; Faraone and Larsson 2019; Rhee and Waldman 2002).
Importantly, while the heritability for each of these individual phenotypes is moderate-to-strong, the genetic variation impacting each of these disorders appears to be largely shared. Each of these phenotypes load onto a single highly heritable (h 2 ~80%) externalizing factor (Kendler et al. 2003b; Krueger et al. 2002; Young et al. 2000), which explains a large proportion of the genetic variance in each individual trait. For example, a general externalizing factor explains 74–80% of genetic influences for AUD, 62–74% for other SUD, and 57–92% for antisocial personality disorder (Kendler and Myers 2013). Other nonclinical risky behaviors that load on to this genetic factor for externalizing include driving while drunk, earlier age at first sex, and riskier sex (Harden et al. 2008b; Quinn and Harden 2013; Samek et al. 2014). Finally, in addition to behaviors, personality traits of novelty seeking, sensation seeking, lack of agreeableness, and lack of conscientiousness also load strongly on this externalizing factor (Kendler and Myers 2013; Krueger et al. 2002; Mann et al. 2015; Young et al. 2000). Overall, twin and family studies indicate that common genetic influences impact multiple traits on the externalizing spectrum.
2.2 Changes in the Etiology of Externalizing Problems Across Development
Like many other complex traits, genetic influences on externalizing problems change across the life course. Genetic influences generally become more important as individuals age and begin to achieve more independence (Dick 2011a; Kendler et al. 2008; Long et al. 2017). This is especially true for traits on the externalizing spectrum. Twin studies repeatedly show that shared environment has important effects on substance use/misuse in early life, whereas genetic influences become more important as individuals reach early adulthood (Dick 2011a; Kendler et al. 2008; Long et al. 2017). Figure 1 provides an overview of the changing relative influence of genetic and shared environmental variance over adolescence (Dick 2011a).
Relative importance of additive genetic (A) and shared environmental (C) influences on alcohol initiation and frequency of use across adolescence (data reported in Dick 2011a). Across adolescence additive genetic influences (A) become more important, while shared environmental influences (C) become less important
While the importance of genetic influences appears to increase over the early life course, there is evidence that the source of genetic influences is relatively stable over time. Multivariate twin models find that the majority of genetic influence on externalizing is attributable to a single factor that explains a large portion of the variance across development (Wichers et al. 2013). Longitudinal models demonstrate that the genetic influences on initial levels of externalizing mostly overlap with the genetic influences on change over time (Hatoum et al. 2018). In other words, the genetic influences that influence externalizing problems in early development also affect behaviors during adolescence. These patterns are similar when we look at specific behaviors on the externalizing spectrum, including alcohol use (Long et al. 2017) and problematic alcohol use (van Beek et al. 2012): the sources of genetic influences are fairly stable across time. Stability in the source of genetic influences over development also occurs with the personality correlates of externalizing problems (Briley and Tucker-Drob 2017). Longitudinal analyses of lab-based tasks related to both impulsivity and delay discounting reveal developmentally stable, genetic influences on these measures meant to assess dimensions of personality (Anokhin et al. 2011; Niv et al. 2012). Overall, it appears that while genetic influences may become more important over time, the same genetic influences act across development.
Despite the evidence that genetic influences on externalizing are stable over development, there is some evidence that the specificity of genetic influences can change for certain phenotypes on the externalizing spectrum. This is especially apparent in regards to alcohol misuse. Genetic risk for broader externalizing problems and genetic risk for AUD both independently predict alcohol misuse across early development. However, the effect size for each form of genetic risk changes over time. In adolescence, broader externalizing risk has a stronger effect on alcohol misuse, while alcohol specific risk becomes important during adulthood (see Fig. 2) (Kendler et al. 2011; Meyers et al. 2014). For other drug use, common genetic influences explain approximately half of the correlation between externalizing problems in childhood and drug initiation in late adolescence (Korhonen et al. 2012).
Genetic risk and alcohol phenotypes from Kendler et al. (2011) and Meyers et al. (2014). Regression coefficients between genetic risk specific to AUD (AUD) or broader externalizing disorders (EXT) and alcohol consumption across age in the Virginia Adult Twin Study of Psychiatric and Substance Use Disorders (VATSPUD, left) and the Finnish Twin Cohort (FinnTwin12, right). Genetic risk for broader externalizing problems and genetic risk specific to AUD both independently predict alcohol misuse across development, though the effect sizes for each changes over time. In adolescence, EXT has a stronger effect on alcohol misuse, while AUD becomes important during adulthood
2.3 Gene-Environment Interactions in Externalizing Problems
Environmental conditions can alter the importance of genetic influences on externalizing behaviors. This phenomenon is referred to as gene-environment interaction, or GxE (Dick 2011b). Researchers have put forth a variety of theoretical models of GxE. For externalizing phenotypes, we see consistent evidence for two primary theoretical paradigms of GxE: the social control/opportunity model and the social distinction model (Boardman et al. 2013; Shanahan and Hofer 2005). Under the social control/opportunity model of GxE, genetic influences become more important under conditions of reduced social control or increased social opportunity (Shanahan and Hofer 2005). For example, environmental conditions related to low social control/increased social opportunity, such as peer deviance (Cooke et al. 2015; Harden et al. 2008a; Mann et al. 2016; Samek et al. 2016) or high neighborhood turnover rate (Dick et al. 2009), are associated with increases in genetic influences on various externalizing traits. On the opposite side, environmental conditions associated with greater social control/reduced social opportunity, such as greater parental monitoring (Cooke et al. 2015; Dick et al. 2007) or involvement in a committed relationship (Barr et al. 2017; Heath et al. 1989), are associated with reduced genetic influences on these phenotypes. It is important to note that this model of GxE spans various externalizing phenotypes including alcohol use, smoking, behavior problems, and delinquency (Cooke et al. 2015; Harden et al. 2008a; Mann et al. 2016; Samek et al. 2016).
Under the social distinction model of GxE (Boardman et al. 2013), certain social conditions “push” the phenotype and increase the importance of environmental influences on a trait at one end of the environmental spectrum. This increase in the importance of environmental variance reduces the importance of genetic influences on that same end of the spectrum. For externalizing problems, childhood socioeconomic status (SES) has consistently fit this model of GxE. Childhood SES moderates the effect of genetic variation on externalizing problems, such that under conditions of lower SES, environmental sources of variance are more important than genetic influences (Middeldorp et al. 2014; Tuvblad et al. 2006). Family SES moderates genetic liability for externalizing whereby higher SES and higher genetic risk are associated with a steeper increase in alcohol problems across adolescence (Barr et al. 2018). Neighborhood-level SES, moderates genetic risk on delinquency (Beaver 2011) and non-violent conduct problems (Burt et al. 2016) in the same direction as family-level SES, such that environmental influences are stronger under conditions of low neighborhood SES. Overall, GxE findings for the social distinction model and social control/opportunity model demonstrate the ways in which the importance genetic influences can shift across environmental conditions.
3 Molecular Genetic Studies of Externalizing Problems
While twin and family data provide valuable insight into the genetics of externalizing problems and other complex traits, they use a latent approach that does not provide information about the specific genetic variants associated with a given trait. Over the past 20 years, the growth in research examining measured genetic variants has rapidly expanded. Much of the early work focused on candidate genes, which were proposed to be associated with a trait because of a hypothesized biological mechanism. Research in this tradition largely focused on genes or single nucleotide polymorphisms (SNPs) in the serotonergic or dopaminergic region. However, candidate gene research has been plagued by false positives, publication bias, and low powered studies (Duncan and Keller 2011). Recent large-scale meta-analyses reveal no support for much of the early work on candidate gene analyses (Border et al. 2019), suggesting that our “best guesses” for genes involved in the underlying biology were not very good. Importantly, candidate gene studies do not fit with our current polygenic understanding of complex traits, whereby phenotypes are influenced by many variants (in the hundreds, if not thousands) of very small effect (Visscher et al. 2017).
With the mapping of the human genome, the focus on candidate gene research has given way to agnostic methods of gene identification that scan the entire genome for SNPs associated with a given trait. Rather than focusing on a single variant with some hypothesized biological mechanism, genome-wide association studies, or GWAS, test the association between a phenotype and SNPs spanning the entire genome. Because of the large number of tests (the typical p-value for genome-wide significance, or GWS, is p < 5 × 10−8 to correct for approximately one million independent tests) and small effect sizes associated with individual variants, adequate sample sizes for discovery GWAS likely require hundreds of thousands to millions of individuals. Fortunately, with the growth of cheaply available genotyping arrays, large-scale biobanks that genotype large numbers of individuals, and direct-to-consumer genetic testing companies that amass genotypic information on large samples, the sample sizes for these GWAS have been rapidly increasing (Mills and Rahal 2019).
3.1 Current GWAS of Externalizing Phenotypes
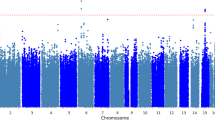
Table 1 provides a sampling of the current GWAS for externalizing traits. To date, these GWAS have predominantly focused on single phenotypes that would be considered part of the externalizing spectrum, mostly substance use outcomes. The majority of GWAS on substance use have focused on alcohol-related phenotypes, including alcohol dependence (Walters et al. 2018) (3 GWS SNPs), alcohol use disorder (Kranzler et al. 2019) (10 GWS SNPs), number of alcoholic drinks per week (Liu et al. 2019) (156 GWS SNPs), and maximum alcohol intake (Gelernter et al. 2019) (6 GWS SNPs). A SNP in the ADH1B gene region (rs1229984) responsible for alcohol metabolism is the most consistently associated SNP across these alcohol GWAS (Gelernter et al. 2019; Kranzler et al. 2019; Liu et al. 2019; Walters et al. 2018); however, other genome-wide significant variants have also begun to emerge, such as those in GCKR which is involved in sugar metabolism in the liver and pancreas (Gelernter et al. 2019; Kranzler et al. 2019; Liu et al. 2019). Sample sizes for these alcohol phenotypes have ranged from moderately to extremely well powered (N’s ~50k – one million). Interestingly, these GWAS reveal that genetic influences on alcohol consumption only partially overlap with variants that impact alcohol-related problems (Sanchez-Roige et al. 2018; Walters et al. 2018).
GWAS of some illicit drugs, especially cannabis phenotypes, are beginning to reach sample sizes that have adequate power for detection of genetic effects. A recent GWAS of lifetime cannabis use (Pasman et al. 2018) (N ~180K) identified eight GWS SNPs. Three of these loci were in the CADM2, which has also shown up in GWAS of impulsivity (Sanchez-Roige et al. 2019), alcohol consumption (Clarke et al. 2017), number of offspring (Day et al. 2016), and risk-taking behavior (Day et al. 2016; Karlsson Linnér et al. 2019). A GWAS of cannabis use disorder (Demontis et al. 2019a) in ~50K individuals identified a single GWS SNP that is a strong expression quantitative trait locus (eQTL, a variant that influences the expression of a gene or genes) for CHRNA2, a nicotine receptor gene related to smoking behavior (Liu et al. 2019). GWAS of other illicit substance use disorders, including cocaine dependence (Gelernter et al. 2014) and opioid dependence (Cheng et al. 2018), are currently underpowered due to extremely small sample sizes (N ~2,500–7,500). Overall, larger sample sizes are needed across illicit substance use to better detect variants associated with these phenotypes.
Beyond substance use, GWAS have focused on other disorders and personality traits related to the externalizing spectrum. A recent GWAS of ADHD (Demontis et al. 2019b) (N ~55K) identified 12 GWS loci. Several of the loci associated with ADHD are located in or near genes implicated in neurodevelopmental processes, including FOXP2, SORCS3, and DUSP6 (Demontis et al. 2019b). For other behavioral phenotypes, a GWAS of antisocial behaviors in ~16k individuals using a broad variety of behaviors (including conduct disorder, behavior check lists, and other scales of antisocial behaviors) did not identify any genome-wide significant loci. GWAS of impulsivity scales including the Barratt Impulsiveness Scale, or BIS (Sanchez-Roige et al. 2019), the composite UPPS-P scale (urgency, premeditation, perseverance, sensation seeking, and positive urgency), and its subscales identified GWAS SNPs in the CADM2 gene region for the sensation seeking subscale of the UPPS-P and CACNA1I (which encodes for a protein thought to be involved in calcium signaling in neurons) gene region for the negative urgency subscale (Sanchez-Roige et al. 2019).
Recently, a GWAS of general risk tolerance in approximately 940K individuals identified 124 independent GWS loci (Karlsson Linnér et al. 2019). This study also examined a composite index of risky behaviors (defined as the first principal component of ever smoking, drinks per week, automobile speeding, and number of sexual partners, N = 315,894) identifying 106 GWS SNPs. The top variants in this GWAS were in the MAPT, CADM2, and FOXP1 gene regions, again implicating genes thought to be involved in neurodevelopmental processes (Karlsson Linnér et al. 2019).
Overall, current gene identification efforts for externalizing traits have begun to detect robust associations with individual disorders/phenotypes on the externalizing spectrum, and more recently, with general externalizing behavior. However, much remains to be discovered as to the biological mechanisms through which these variants influence behavior. While some variants have well-known biological function (such as alcohol metabolism or nicotine receptor genes), others (such as those related to neurodevelopmental processes and brain function) need further scrutiny. Future research will need to move beyond simple associations. Integrating data from human GWAS into model organisms will allow us to directly test the biological function of genes identified in GWAS and whether or not these genes exert some causal influence on externalizing problems (Baker et al. 2011; Jay 2012). As the cost of whole genome sequencing comes down, we will also be better able to examine the impact of rare variants, which are largely excluded from current methods (which focus primarily on common variants). Finally, as we begin to think of genes and variants as parts of dense networks, we may better understand the underlying biological mechanisms between genotype and phenotype (Visscher et al. 2017).
3.2 Genetic Correlations and Multivariate Genomic Methods
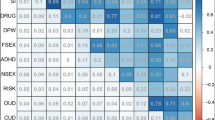
Perhaps the most interesting finding to emerge from all of these GWAS of phenotypes on the externalizing spectrum is the strong genetic overlap between traits, confirming earlier results from twin and family studies. Alcohol use disorder, alcohol consumption, smoking status, lifetime cannabis use, risky behaviors, general risk tolerance, and polysubstance use all have significant genetic correlations with one another (Kranzler et al. 2019; Liu et al. 2019; Meyers et al. 2014; Sanchez-Roige et al. 2019; Walters et al. 2018). These externalizing phenotypes also overlap genetically with other socio-demographic outcomes related to externalizing, including age at first birth, number of children, and educational attainment (Kranzler et al. 2019; Sanchez-Roige et al. 2019; Walters et al. 2018). Figure 3 Footnote 2 shows the genetic correlations between a subset of externalizing phenotypes, estimated using GWAS summary statistics (Demontis et al. 2019b; Karlsson Linnér et al. 2019; Liu et al. 2019; Pasman et al. 2018; Sanchez-Roige et al. 2018; Walters et al. 2018). Genetic correlations were calculated using bivariate LD score regression (Bulik-Sullivan et al. 2015).
Genetic correlations calculated from published GWAS of externalizing phenotypes (using bivariate LD score regression). Heatmap of genetic correlation using effect sizes from currently published GWAS, with correlation estimates denoted in the cells. There is a strong pattern of significant genetic overlap between clinically relevant phenotypes (Problematic Alcohol Use, ADHD), other substance use (Lifetime Cannabis Use, Ever Smoker), risky sexual behaviors (Age at First Sex, Number of Sexual Partners), and personality (General Risk Tolerance). The results in the heatmap demonstrate a potential shared genetic influence across these phenotypes. All genetic correlations are significant after correcting for multiple testing ( p < 0.0024)
There are now concerted efforts to use information from these and other GWAS to move beyond univariate analyses and model the multivariate genetic architecture of externalizing problems identified in twin and family studies. One such ongoing project is the Externalizing Consortium (Dick et al. 2018). New multivariate gene identification methods such as Genomic Structural Equation Modeling, or Genomic SEM (Grotzinger et al. 2019), utilize genetic correlations to model the underlying factor structure of a set of phenotypes using GWAS summary statistics. While traditional SEM models the phenotypic covariance to measure a latent factor, Genomic SEM models a latent genetic factor based on the genetic covariance. Utilizing these new multivariate methods allows one to boost power to identify genetic variants by harnessing existing GWAS of genetically correlated phenotypes. This type of multivariate analysis illustrates the advantage of combining information across externalizing traits to detect genetic variants associated with a range of externalizing outcomes. As more well-powered GWAS of externalizing phenotypes become available, we will be able to model the underlying externalizing spectrum with even more power and precision.
3.3 Research Using PRS for Externalizing Phenotypes
Beyond identifying associations between individual variants and a phenotype, GWAS results can be used to create polygenic risk scores (PRS) that index an individual’s overall liability for the outcomes, in order to study associations between these aggregate measures of genetic risk and phenotypes in external samples. An important component of using PRS is that the sample in which they are used must be independent of the discovery GWAS sample. Figure 4 provides an overview of how PRS are constructed. PRS are computed as the average of the number of “risk” alleles that an individual carries weighted by the parameter estimates (e.g., betas, odds ratios, and Z-scores) identified in a GWAS. Because SNPs that are close to one another in the genome correlated (referred to as linkage disequilibrium, or LD), PRS are generally constructed from a subset of independent SNPs. These SNPs can be selected using a variety of methods including “pruning and thresholding,” where SNPs below a certain GWAS p-value and LD threshold (using r 2) are included (International Schizophrenia Consortium 2009); or LDpred, which uses a Bayesian approach to model SNP effect sizes while accounting for LD from an external reference panel (Vilhjalmsson et al. 2015).
Hypothetical example for calculating polygenic risk scores. Example of using GWAS summary statistics (left) to calculate polygenic risk scores (PRS) in an independent sample. GWAS provides the effect sizes (Beta) and Risk allele to calculate the weighted sum of risk alleles that an individual in the target sample carries. For example, Person 1 carries two risk alleles (A) at SNP 1, a single risk allele (T) at SNP 2, and a single risk allele (G) at SNP 3. Therefore, there PRS would be 2∗0.874 + 1∗−0.007 + 1∗0.148 = 1.889. This process occurs across all SNPs included for calculating a given PRS for each person in the independent target sample
PRS provide a flexible way of taking results from large-scale GWAS into samples with extensive phenotyping or longitudinal data to answer more nuanced questions about how genetic liability unfolds over time or how it changes across the specific environments. Extending the twin-family literature, research that uses PRS for externalizing problems has also found evidence of GxE. Following the social control/social opportunity model of GxE from the twin literature, recent work using PRS has found evidence of romantic partnerships moderating the association between PRS and alcohol misuse (Barr et al. 2019); peer deviance and parental monitoring moderating the association between PRS and externalizing disorders (Salvatore et al. 2014); and neighborhood social cohesion moderating the association between PRS and nicotine use (Meyers et al. 2013). In each of the listed PRS analyses, the association between the PRS and the corresponding phenotype was stronger under conditions of reduced social control (e.g., not in a relationship, association with deviant peers, low parental monitoring, and low neighborhood social cohesion) compared to the conditions of increased control/reduced opportunity.
PRS derived from GWAS of other outcomes that are genetically correlated to externalizing problems can also be used as proxies for PRS of externalizing problems. For example, PRS derived from a GWAS of educational attainment (Lee et al. 2018) predict antisocial behaviors across the life course, from early adolescence into adulthood (Wertz et al. 2018). PRS derived from GWAS of schizophrenia (Schizophrenia Working Group of the Psychiatric Genomics Consortium 2014) also predict childhood behavior problems (Jansen et al. 2018). Finally, PRS derived from a GWAS of educational attainment predict a variety of substance use disorders, including alcohol, tobacco, and cannabis use disorders (Salvatore et al. 2019). These analyses demonstrate that in the absence of well-powered GWAS of externalizing problems, PRS derived from large GWAS of genetically correlated phenotypes provide information as to how these forms of genetic liability unfold over time and relate to multiple externalizing phenotypes.
It is important to note that even though PRS have proven to be a useful tool for understanding genetic liability, even in large cohorts, PRS continue to predict only small portions of the variance in independent samples. For example, PRS from a recent GWAS approximately one million explained ~2.5% of variance in alcohol consumption (Liu et al. 2019). Additionally, PRS aggregate information from across the genome without regards to the biological function of the variants included. Future methods that can incorporate additional information from biological annotations or functional enrichment may improve our ability to predict these disorders from PRS (Márquez-Luna et al. 2019).
3.4 Increasing Diversity in Genetic Research
Concerted efforts to increase the diversity of participants in genomic research are vital. To date, large scale GWAS are composed almost entirely of individuals of European ancestry (Mills and Rahal 2019). Inclusion of individuals of diverse ancestries is scientifically important because including diverse ancestries increases the discovery power in GWAS (Dick et al. 2017; Wojcik et al. 2019) and differences in LD structure across allow us to get closer to causal variants (Bigdeli et al. 2019). Creating more diverse samples is also important for ethical reasons. PRS derived from ancestral populations that differ from the target sample perform poorly (Martin et al. 2017). In the push towards precision medicine, the current GWAS will likely exacerbate health disparities rather than help solve them (Martin et al. 2019). Therefore, greater diversity in genomic research is both a moral and scientific imperative.
4 Conclusion
Research into the etiology of externalizing problems has found that each of the psychiatric disorders, nonclinical behaviors, and personality characteristics on the externalizing spectrum are heritable to varying degrees. Overall, these externalizing problems share a common genetic etiology that accounts for a large share of the genetic variance in each of the corresponding phenotypes. However, the importance of these genetic influences can change across developmental and environmental context, typically becoming more influential as individuals reach young adulthood and under conditions in which social control is limited. Recent work has moved beyond the latent genetic designs of twin and family research into designs using measure genome-wide data. While we have begun to robustly detect variants associated with these traits, there is still considerable work to be done on elucidating biological mechanisms of risk. Future multivariate GWAS efforts will help to better understand the underlying genetic architecture shared across individual phenotypes. As we better understand the genetic architecture, we will be able to use results to create more powerful PRS in independent samples to further explore the ways in which risk unfolds across time and environmental context. And as we move towards the era of precision medicine, we will need to ensure that we have even larger discovery samples of diverse ancestries so that our results are able to be used to improve the health of all individuals in the population.
Notes
- 1.
Heritability (h 2) generally refers to broad sense heritability, or the proportion of variance in a population that is the result of genetic influences. Twin and family models generally divide variance in a phenotype into additive genetic (A), shared environmental (C), and unique environmental (E) variance. Unique environmental influences also include measurement error.
- 2.
We would like to thank Richard Karlsson-Linnér for his work gathering these GWAS summary statistics, cleaning the data, and running bivariate LDSC.
References
Anokhin AP, Golosheykin S, Grant JD, Heath AC (2011) Heritability of delay discounting in adolescence: a longitudinal twin study. Behav Genet 41:175–183. https://doi.org/10.1007/s10519-010-9384-7
Baker EJ, Jay JJ, Bubier JA, Langston MA, Chesler EJ (2011) GeneWeaver: a web-based system for integrative functional genomics. Nucleic Acids Res 40:D1067–D1076. https://doi.org/10.1093/nar/gkr968
Barr PB et al (2017) Social relationships moderate genetic influences on heavy drinking in Young adulthood. J Stud Alcohol Drugs 78:817–826
Barr PB, Silberg J, Dick DM, Maes HH (2018) Childhood socioeconomic status and longitudinal patterns of alcohol problems: variation across etiological pathways in genetic risk. Soc Sci Med 209:51–58. https://doi.org/10.1016/j.socscimed.2018.05.027
Barr PB et al (2019) Polygenic risk for alcohol misuse is moderated by romantic partnerships. Addiction 114:1753. https://doi.org/10.1111/add.14712
Beaver KM (2011) Environmental moderators of genetic influences on adolescent delinquent involvement and victimization. J Adolesc Res 26:84–114. https://doi.org/10.1177/0743558410384736
Bigdeli TB et al (2019) Contributions of common genetic variants to risk of schizophrenia among individuals of African and Latino ancestry. Mol Psychiatry. https://doi.org/10.1038/s41380-019-0517-y
Boardman JD, Daw J, Freese J (2013) Defining the environment in gene--environment research: lessons from social epidemiology. Am J Public Health 103:S64–S72. https://doi.org/10.2105/AJPH.2013.301355
Border R, Johnson EC, Evans LM, Smolen A, Berley N, Sullivan PF, Keller MC (2019) No support for historical candidate gene or candidate gene-by-interaction hypotheses for major depression across multiple large samples. Am J Psychiatry 176:376–387. https://doi.org/10.1176/appi.ajp.2018.18070881
Briley DA, Tucker-Drob EM (2017) Comparing the developmental genetics of cognition and personality over the life span. J Pers 85:51–64. https://doi.org/10.1111/jopy.12186
Bulik-Sullivan BK et al (2015) LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet 47:291–295. https://doi.org/10.1038/ng.3211
Burt SA (2009) Are there meaningful etiological differences within antisocial behavior? Results of a meta-analysis. Clin Psychol Rev 29:163–178. https://doi.org/10.1016/j.cpr.2008.12.004
Burt SA, Klump KL, Gorman-Smith D, Neiderhiser JM (2016) Neighborhood disadvantage alters the origins of Children’s nonaggressive conduct problems. Clin Psychol Sci 4:511–526. https://doi.org/10.1177/2167702615618164
Center for Disease Control and Prevention (2019) Mortality Data—National Vital Statistics System. https://www.cdc.gov/nchs/nvss/deaths.htm
Chen P, Jacobson KC (2012) Developmental trajectories of substance use from early adolescence to young adulthood: gender and racial/ethnic differences. J Adolesc Health 50:154–163. https://doi.org/10.1016/j.jadohealth.2011.05.013
Cheng Z, Zhou H, Sherva R, Farrer LA, Kranzler HR, Gelernter J (2018) Genome-wide association study identifies a regulatory variant of RGMA associated with opioid dependence in European Americans. Biol Psychiatry 84:762–770. https://doi.org/10.1016/j.biopsych.2017.12.016
Clarke TK et al (2017) Genome-wide association study of alcohol consumption and genetic overlap with other health-related traits in UK Biobank (N=112 117). Mol Psychiatry 22:1376. https://doi.org/10.1038/mp.2017.153. https://www.nature.com/articles/mp2017153#supplementary-information
Cooke ME et al (2015) Gene-environment interaction effects of peer deviance, parental knowledge and stressful life events on adolescent alcohol use. Twin Res Hum Genet 18:507–517. https://doi.org/10.1017/thg.2015.56
Day FR et al (2016) Physical and neurobehavioral determinants of reproductive onset and success. Nat Genet 48:617. https://doi.org/10.1038/ng.3551. https://www.nature.com/articles/ng.3551#supplementary-information
Degenhardt L et al (2013) The global epidemiology and contribution of Cannabis use and dependence to the global burden of disease: results from the GBD 2010 study. PLoS One 8:e76635. https://doi.org/10.1371/journal.pone.0076635
Demontis D et al (2019a) Genome-wide association study implicates CHRNA2 in cannabis use disorder. Nat Neurosci 22:1066–1074. https://doi.org/10.1038/s41593-019-0416-1
Demontis D et al (2019b) Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat Genet 51:63–75. https://doi.org/10.1038/s41588-018-0269-7
Dick DM (2011a) Developmental changes in genetic influences on alcohol use and dependence. Child Dev Perspect 5:223–230. https://doi.org/10.1111/j.1750-8606.2011.00207.x
Dick DM (2011b) Gene-environment interaction in psychological traits and disorders. Annu Rev Clin Psychol 7:383–409. https://doi.org/10.1146/annurev-clinpsy-032210-104518
Dick DM, Viken R, Purcell S, Kaprio J, Pulkkinen L, Rose RJ (2007) Parental monitoring moderates the importance of genetic and environmental influences on adolescent smoking. J Abnorm Psychol 116:213–218. https://doi.org/10.1037/0021-843X.116.1.213
Dick DM, Bernard M, Aliev F, Viken R, Pulkkinen L, Kaprio J, Rose RJ (2009) The role of socioregional factors in moderating genetic influences on early adolescent behavior problems and alcohol use. Alcohol Clin Exp Res 33:1739–1748. https://doi.org/10.1111/j.1530-0277.2009.01011.x
Dick DM, Barr P, Guy M, Nasim A, Scott D (2017) Review: genetic research on alcohol use outcomes in African American populations: a review of the literature, associated challenges, and implications. Am J Addict 26:486–493. https://doi.org/10.1111/ajad.12495
Dick DM et al (2018) Using the genetic architecture of externalizing disorders to aid in gene identification. Paper presented at the annual meeting of the behavior genetics association, Boston, MA
Duncan LE, Keller MC (2011) A critical review of the first 10 years of candidate gene-by-environment interaction research in psychiatry. Am J Psychiatry 168:1041–1049. https://doi.org/10.1176/appi.ajp.2011.11020191
Faraone SV, Larsson H (2019) Genetics of attention deficit hyperactivity disorder. Mol Psychiatry 24:562–575. https://doi.org/10.1038/s41380-018-0070-0
Florence CS, Zhou C, Luo F, Xu L (2016) The economic burden of prescription opioid overdose, abuse, and dependence in the United States, 2013. Med Care 54:901–906. https://doi.org/10.1097/MLR.0000000000000625
Gelernter J, Sherva R, Koesterer R, Almasy L, Zhao H, Kranzler HR, Farrer L (2014) Genome-wide association study of cocaine dependence and related traits: FAM53B identified as a risk gene. Mol Psychiatry 19:717–723. https://doi.org/10.1038/mp.2013.99
Gelernter J et al (2019) Genomewide Association Study of Maximum Habitual Alcohol Intake in >140,000 US European-and African-American Veterans Yields Novel Risk Loci Biological Psychiatry
Grant BF et al (2015) Epidemiology of DSM-5 alcohol use disorder: results from the National Epidemiologic Survey on alcohol and related conditions III. JAMA Psychiat 72:757–766. https://doi.org/10.1001/jamapsychiatry.2015.0584
Grant BF et al (2016) Epidemiology of DSM-5 drug use disorder: results from the National Epidemiologic Survey on alcohol and related conditions–III. JAMA Psychiat 73:39–47. https://doi.org/10.1001/jamapsychiatry.2015.2132
Grotzinger AD et al (2019) Genomic structural equation modelling provides insights into the multivariate genetic architecture of complex traits. Nat Hum Behav 3:513. https://doi.org/10.1038/s41562-019-0566-x
Harden KP, Hill JE, Turkheimer E, Emery RE (2008a) Gene-environment correlation and interaction in peer effects on adolescent alcohol and tobacco use. Behav Genet 38:339–347. https://doi.org/10.1007/s10519-008-9202-7
Harden KP, Mendle J, Hill JE, Turkheimer E, Emery RE (2008b) Rethinking timing of first sex and delinquency. J Youth Adolesc 37:373–385. https://doi.org/10.1007/s10964-007-9228-9
Hatoum AS, Rhee SH, Corley RP, Hewitt JK, Friedman NP (2018) Etiology of stability and growth of internalizing and externalizing behavior problems across childhood and adolescence. Behav Genet 48:298. https://doi.org/10.1007/s10519-018-9900-8
Heath AC, Jardine R, Martin NG (1989) Interactive effects of genotype and social environment on alcohol consumption in female twins. J Stud Alcohol 50:38–48
International Schizophrenia Consortium (2009) Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460:748–752. https://doi.org/10.1038/nature08185
Jansen PR et al (2018) Polygenic scores for schizophrenia and educational attainment are associated with behavioural problems in early childhood in the general population. J Child Psychol Psychiatry 59:39–47. https://doi.org/10.1111/jcpp.12759
Jay JJ (2012) Chapter one – cross species integration of functional genomics experiments. In: Chesler EJ, Haendel MA (eds) International review of neurobiology, vol 104. Academic Press, London, pp 1–24. https://doi.org/10.1016/B978-0-12-398323-7.00001-X
Karlsson Linnér R et al (2019) Genome-wide association analyses of risk tolerance and risky behaviors in over 1 million individuals identify hundreds of loci and shared genetic influences. Nat Genet 51:245–257. https://doi.org/10.1038/s41588-018-0309-3
Kendler KS, Myers J (2013) The boundaries of the internalizing and externalizing genetic spectra in men and women. Psychol Med 44:647–655. https://doi.org/10.1017/S0033291713000585
Kendler KS, Karkowski LM, Neale MC, Prescott CA (2000) Illicit psychoactive substance use, heavy use, abuse, and dependence in a US population-based sample of male twins. JAMA Psychiat 57:261–269. https://doi.org/10.1001/archpsyc.57.3.261
Kendler KS, Jacobson KC, Prescott CA, Neale MC (2003a) Specificity of genetic and environmental risk factors for use and abuse/dependence of Cannabis, cocaine, hallucinogens, sedatives, stimulants, and opiates in male twins. Am J Psychiatry 160:687–695. https://doi.org/10.1176/appi.ajp.160.4.687
Kendler KS, Prescott CA, Myers J, Neale MC (2003b) The structure of genetic and environmental risk factors for common psychiatric and substance use disorders in men and women. Arch Gen Psychiatry 60:929–937. https://doi.org/10.1001/archpsyc.60.9.929
Kendler KS, Schmitt E, Aggen SH, Prescott CA (2008) Genetic and environmental influences on alcohol, caffeine, cannabis, and nicotine use from early adolescence to middle adulthood. Arch Gen Psychiatry 65:674–682. https://doi.org/10.1001/archpsyc.65.6.674
Kendler KS, Gardner C, Dick DM (2011) Predicting alcohol consumption in adolescence from alcohol-specific and general externalizing genetic risk factors, key environmental exposures and their interaction. Psychol Med 41:1507–1516. https://doi.org/10.1017/S003329171000190X
Kessler RC, Berglund P, Demler O, Jin R, Merikangas KR, Walters EE (2005a) Lifetime prevalence and age-of-onset distributions of DSM-IV disorders in the national comorbidity survey replication. Arch Gen Psychiatry 62:593–602. https://doi.org/10.1001/archpsyc.62.6.593
Kessler RC, Chiu WT, Demler O, Walters EE (2005b) Prevalence, severity, and comorbidity of twelve-month DSM-IV disorders in the National Comorbidity Survey Replication (NCS-R). Arch Gen Psychiatry 62:617–627. https://doi.org/10.1001/archpsyc.62.6.617
Korhonen T, Latvala A, Dick DM, Pulkkinen L, Rose RJ, Kaprio J, Huizink AC (2012) Genetic and environmental influences underlying externalizing behaviors, cigarette smoking and illicit drug use across adolescence. Behav Genet 42:614–625. https://doi.org/10.1007/s10519-012-9528-z
Kranzler HR et al (2019) Genome-wide association study of alcohol consumption and use disorder in 274,424 individuals from multiple populations. Nat Commun 10:1499. https://doi.org/10.1038/s41467-019-09480-8
Krueger RF, Hicks BM, Patrick CJ, Carlson SR, Iacono WG, McGue M (2002) Etiological connections among substance dependence, antisocial behavior and personality: modeling the externalizing spectrum. J Abnorm Psychol 111:411–424. https://doi.org/10.1037//0021-843x.111.3.411
Lee JJ et al (2018) Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat Genet 50:1112–1121. https://doi.org/10.1038/s41588-018-0147-3
Liu M et al (2019) Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat Genet 51:237–244. https://doi.org/10.1038/s41588-018-0307-5
Long EC, Verhulst B, Aggen SH, Kendler KS, Gillespie NA (2017) Contributions of genes and environment to developmental change in alcohol use. Behav Genet 47:498–506. https://doi.org/10.1007/s10519-017-9858-y
Maes HH, Sullivan PF, Bulik CM, Neale MC, Prescott CA, Eaves LJ, Kendler KS (2004) A twin study of genetic and environmental influences on tobacco initiation, regular tobacco use and nicotine dependence. Psychol Med 34:1251–1261. https://doi.org/10.1017/S0033291704002405
Mann FD, Briley DA, Tucker-Drob EM, Harden KP (2015) A behavioral genetic analysis of callous-unemotional traits and big five personality in adolescence. J Abnorm Psychol 124:982–993. https://doi.org/10.1037/abn0000099
Mann FD, Patterson MW, Grotzinger AD, Kretsch N, Tackett JL, Tucker-Drob EM, Harden KP (2016) Sensation seeking, peer deviance, and genetic influences on adolescent delinquency: evidence for person-environment correlation and interaction. J Abnorm Psychol 125:679–691. https://doi.org/10.1037/abn0000160
Márquez-Luna C, Gazal S, Loh P-R, Kim SS, Furlotte N, Auton A, Price AL (2019) Modeling functional enrichment improves polygenic prediction accuracy in UK Biobank and 23andMe data sets. bioRxiv:375337. https://doi.org/10.1101/375337
Martin AR et al (2017) Human demographic history impacts genetic risk prediction across diverse populations. Am J Hum Genet 100:635–649. https://doi.org/10.1016/j.ajhg.2017.03.004
Martin AR, Kanai M, Kamatani Y, Okada Y, Neale BM, Daly MJ (2019) Clinical use of current polygenic risk scores may exacerbate health disparities. Nat Genet 51:584–591. https://doi.org/10.1038/s41588-019-0379-x
Maurer D (2017) COSTS OF CRIME: experts report challenges estimating costs and suggest improvements to better inform policy decisions. Washington, DC
Meyers JL et al (2013) Interaction between polygenic risk for cigarette use and environmental exposures in the Detroit neighborhood health study. Transl Psychiatry 3:e290. https://doi.org/10.1038/tp.2013.63
Meyers JL et al (2014) Genetic influences on alcohol use behaviors have diverging developmental trajectories: a prospective study among male and female twins. Alcohol Clin Exp Res 38:2869–2877. https://doi.org/10.1111/acer.12560
Middeldorp CM, Lamb DJ, Vink JM, Bartels M, van Beijsterveldt CE, Boomsma DI (2014) Child care, socio-economic status and problem behavior: a study of gene-environment interaction in young Dutch twins. Behav Genet 44:314–325. https://doi.org/10.1007/s10519-014-9660-z
Mills MC, Rahal C (2019) A scientometric review of genome-wide association studies. Commun Biol 2:9. https://doi.org/10.1038/s42003-018-0261-x
National Drug Intelligence Center (2011) National Drug Threat Assessment. www.justice.gov/archive/ndic/pubs44/44849/44849p.pdf
Neale MC, Cardon LR (2013) Methodology for genetic studies of twins and families, vol 67. Springer Science & Business Media, New York
Niv S, Tuvblad C, Raine A, Wang P, Baker LA (2012) Heritability and longitudinal stability of impulsivity in adolescence. Behav Genet 42:378–392. https://doi.org/10.1007/s10519-011-9518-6
Pasman JA et al (2018) GWAS of lifetime cannabis use reveals new risk loci, genetic overlap with psychiatric traits, and a causal influence of schizophrenia. Nat Neurosci 21:1161–1170. https://doi.org/10.1038/s41593-018-0206-1
Polanczyk G, de Lima MS, Horta BL, Biederman J, Rohde LA (2007) The worldwide prevalence of ADHD: a systematic review and metaregression analysis. Am J Psychiatry 164:942–948. https://doi.org/10.1176/ajp.2007.164.6.942
Powell D, Perreira KM, Harris KM (2010) Trajectories of delinquency from adolescence to adulthood. Youth Soc 41:475–502
Quinn PD, Harden KP (2013) Behind the wheel and on the map: genetic and environmental associations between drunk driving and other externalizing behaviors. J Abnorm Psychol 122:1166–1178. https://doi.org/10.1037/a0034426.supp
Reitsma MB et al (2017) Smoking prevalence and attributable disease burden in 195 countries and territories, 1990–2015: a systematic analysis from the global burden of disease study 2015. Lancet 389:1885–1906. https://doi.org/10.1016/S0140-6736(17)30819-X
Rhee SH, Waldman ID (2002) Genetic and environmental influences on antisocial behavior: a meta-analysis of twin and adoption studies. Psychol Bull 128:490–529. https://doi.org/10.1037/0033-2909.128.3.490
Sacks JJ, Gonzales KR, Bouchery EE, Tomedi LE, Brewer RD (2015) 2010 national and state costs of excessive alcohol consumption. Am J Prev Med 49:e73–e79. https://doi.org/10.1016/j.amepre.2015.05.031
Salvatore JE, Dick DM (2018) Genetic influences on conduct disorder. Neurosci Biobehav Rev 91:91–101. https://doi.org/10.1016/j.neubiorev.2016.06.034
Salvatore JE et al (2014) Polygenic risk for externalizing disorders: gene-by-development and gene-by-environment effects in adolescents and Young adults. Clin Psychol Sci 3:189–201. https://doi.org/10.1177/2167702614534211
Salvatore JE et al (2019) Sibling comparisons elucidate the associations between educational attaniment polygenic scores and alcohol, nicotine, and cannabis. Addiction. https://doi.org/10.1111/add.14815
Samek DR, Iacono WG, Keyes MA, Epstein M, Bornovalova MA, McGue M (2014) The developmental progression of age 14 behavioral disinhibition, early age of sexual initiation, and subsequent sexual risk-taking behavior. J Child Psychol Psychiatry 55:784–792. https://doi.org/10.1111/jcpp.12176
Samek DR, Hicks BM, Keyes MA, Iacono WG, McGue M (2016) Antisocial peer affiliation and externalizing disorders: evidence for gene × environment × development interaction. Dev Psychopathol 29:155–172. https://doi.org/10.1017/S0954579416000109
Sanchez-Roige S et al (2018) Genome-wide association study meta-analysis of the Alcohol Use Disorders Identification Test (AUDIT) in two population-based cohorts. Am J Psychiatry. https://doi.org/10.1176/appi.ajp.2018.18040369
Sanchez-Roige S, Fontanillas P, Elson SL, Gray JC, de Wit H, MacKillop J, Palmer AA (2019) Genome-wide association studies of impulsive personality traits (BIS-11 and UPPS-P) and drug experimentation in up to 22,861 adult research participants identify loci in the CACNA1I and CADM2 genes. J Neurosci 39:2562. https://doi.org/10.1523/JNEUROSCI.2662-18.2019
Schizophrenia Working Group of the Psychiatric Genomics Consortium (2014) Biological insights from 108 schizophrenia-associated genetic loci. Nature 511:421. https://doi.org/10.1038/nature13595. https://www.nature.com/articles/nature13595#supplementary-information
Shanahan MJ, Hofer SM (2005) Social context in gene-environment interactions: retrospect and prospect. J Gerontol B Psychol Sci Soc Sci 60:65–76
Tielbeek JJ, Johansson A, Polderman TJ, Rautiainen MR, Jansen P, Taylor M et al (2017) Genome-wide association studies of a broad spectrum of antisocial behavior. JAMA Psychiatry 74(12):1242–1250
Tuvblad C, Grann M, Lichtenstein P (2006) Heritability for adolescent antisocial behavior differs with socioeconomic status: gene-environment interaction. J Child Psychol Psychiatry 47:734–743. https://doi.org/10.1111/j.1469-7610.2005.01552.x
van Beek JHDA et al (2012) Stable genetic effects on symptoms of alcohol abuse and dependence from adolescence into early adulthood. Behav Genet 42:40–56. https://doi.org/10.1007/s10519-011-9488-8
van den Bree MBM, Johnson EO, Neale MC, Pickens RW (1998) Genetic and environmental influences on drug use and abuse/dependence in male and female twins. Drug Alcohol Depend 52:231–241. https://doi.org/10.1016/S0376-8716(98)00101-X
Verhulst B, Neale MC, Kendler KS (2015) The heritability of alcohol use disorders: a meta-analysis of twin and adoption studies. Psychol Med 45:1061–1072. https://doi.org/10.1017/S0033291714002165
Verweij KJ et al (2010) Genetic and environmental influences on cannabis use initiation and problematic use: a meta-analysis of twin studies. Addiction 105:417–430. https://doi.org/10.1111/j.1360-0443.2009.02831.x
Vilhjalmsson BJ et al (2015) Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am J Hum Genet 97:576–592. https://doi.org/10.1016/j.ajhg.2015.09.001
Visscher PM, Wray NR, Zhang Q, Sklar P, McCarthy MI, Brown MA, Yang J (2017) 10 years of GWAS discovery: biology, function, and translation. Am J Hum Genet 101:5–22. https://doi.org/10.1016/j.ajhg.2017.06.005
Walters RK et al (2018) Transancestral GWAS of alcohol dependence reveals common genetic underpinnings with psychiatric disorders. Nat Neurosci 21:1656–1669. https://doi.org/10.1038/s41593-018-0275-1
Wertz J et al (2018) Genetics and crime: integrating new genomic discoveries into psychological research about antisocial behavior. Psychol Sci 29:791–803. https://doi.org/10.1177/0956797617744542
Wichers M, Gardner C, Maes HH, Lichtenstein P, Larsson H, Kendler KS (2013) Genetic innovation and stability in externalizing problem behavior across development: a multi-informant twin study. Behav Genet 43:191–201. https://doi.org/10.1007/s10519-013-9586-x
Wojcik GL et al (2019) Genetic analyses of diverse populations improves discovery for complex traits. Nature 570:514–518. https://doi.org/10.1038/s41586-019-1310-4
Young SE, Stallings MC, Corley RP, Krauter KS, Hewitt JK (2000) Genetic and environmental influences on behavioral disinhibition. Am J Med Genet 96:684–695. https://doi.org/10.1002/1096-8628(20001009)96:5<684::aid-ajmg16>3.0.co;2-g
Acknowledgments
We would like to thank our collaborators in the Externalizing Consortium: Richard Karlsson Linnér, Travis Mallard, Sandra Sanchez-Roige, Irwin Waldman, Abraham Palmer, Paige Harden, and Philipp Koellinger. Research reported in this publication was supported by the National Institute on Alcohol Abuse and Alcoholism of the National Institutes of Health under award numbers R01AA015416 and K02AA018755 to DMD. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. This research also used summary data from the Psychiatric Genomics Consortium (PGC) Substance Use Disorders (SUD) working group. The PGC-SUD is supported by funds from NIDA and NIMH to MH109532 and, previously, had analyst support from NIAAA to U01AA008401 (COGA). PGC-SUD gratefully acknowledges its contributing studies and the participants in those studies, without whom this effort would not be possible.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Barr, P.B., Dick, D.M. (2019). The Genetics of Externalizing Problems. In: de Wit, H., Jentsch, J.D. (eds) Recent Advances in Research on Impulsivity and Impulsive Behaviors. Current Topics in Behavioral Neurosciences, vol 47. Springer, Cham. https://doi.org/10.1007/7854_2019_120
Download citation
DOI: https://doi.org/10.1007/7854_2019_120
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-60510-0
Online ISBN: 978-3-030-60511-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)