Abstract
Terpenoids (isoprenoids) represent the largest and most diverse class of chemicals among the myriad compounds produced by plants. Plants employ terpenoid metabolites for a variety of basic functions in growth and development but use the majority of terpenoids for more specialized chemical interactions and protection in the abiotic and biotic environment. Traditionally, plant-based terpenoids have been used by humans in the food, pharmaceutical, and chemical industries, and more recently have been exploited in the development of biofuel products. Genomic resources and emerging tools in synthetic biology facilitate the metabolic engineering of high-value terpenoid products in plants and microbes. Moreover, the ecological importance of terpenoids has gained increased attention to develop strategies for sustainable pest control and abiotic stress protection. Together, these efforts require a continuous growth in knowledge of the complex metabolic and molecular regulatory networks in terpenoid biosynthesis. This chapter gives an overview and highlights recent advances in our understanding of the organization, regulation, and diversification of core and specialized terpenoid metabolic pathways, and addresses the most important functions of volatile and nonvolatile terpenoid specialized metabolites in plants.
Graphical Abstract

Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Introductory chapters on terpenoid biosynthesis usually highlight the large number of terpenoid compounds found in nature. Indeed, the structural diversity associated with at least 40,000 compounds makes the class of terpenoids one of the most impressive examples in the divergent evolution of plant chemicals. The evolutionary success of this compound class is in part based on the simplicity of constructing different size molecules. According to the isoprene rule recognized by Wallach and Rutzicka in the late nineteenth and mid-twentieth centuries [1], all terpenoids are derived from the universal five-carbon building blocks, isopentenyl diphosphate (IPP) and its allylic isomer dimethylallyl diphosphate (DMAPP). The prenyl diphosphate intermediates built by condensation of these five-carbon units are used as precursors for the biosynthesis of terpenoids with fundamental functions in growth and development and for the formation of a large number of terpenoid compounds with more specialized roles in the interaction of plants with their environment. It is the latter group of terpenoids that is characterized by its tremendous structural diversity as a consequence of divergent biosynthetic gene evolution. Specialized terpenoids have a long history of being used as flavors, fragrances, pharmaceuticals, insecticides, and industrial compounds, several of which are addressed in this book. With the growing need for sustainable production platforms of plant-based drugs and the emerging use of terpenoids in the production of alternative fuels, substantial progress has been made in the engineering of terpenoid biosynthetic pathways in microbes and plants [2, 3]. Advanced functional genomics approaches provide unlimited access to the biosynthetic genes and molecular regulators of terpenoid-producing plants, and, at the same time, allow deeper insight to the complexity of plant terpenoid metabolism and regulation. In this chapter, I provide an overview of the organization of the early and core terpenoid metabolic pathways and give updates on the regulation and functional diversification of their genes and enzymes. Furthermore, I summarize the function of terpene synthases and describe aspects of their coordinated and tissue-specific regulation in specialized metabolism prior to addressing the diverse roles of terpenoids in plant–environment interactions.
2 Core Terpenoid Biosynthetic Pathways and Their Regulation
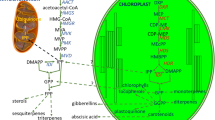
Successful engineering of terpenoid products in plants critically depends on the flux of precursors delivered by the core isoprenoid biosynthetic pathways and, consequently, on the dynamic regulation of these biosynthetic routes. Plants use two independent pathways to produce IPP and DMAPP: the primarily cytosolic mevalonic acid (MVA) pathway and the plastidial methylerythritol phosphate (MEP) pathway (Fig. 1). The MVA pathway predominantly provides the precursors for the cytosolic biosynthesis of sesquiterpenoids, polyprenols, phytosterols, brassinosteroids, and triterpenoids, and for terpenoid biosynthesis in mitochondria (e.g., ubiquinones, polyprenols), and the five-carbon units derived from the MEP pathway are preferably used for the biosynthesis of hemiterpenoids (e.g., isoprene), monoterpenoids, diterpenoids, carotenoids and their breakdown products, cytokinins, gibberellins, chlorophyll, tocopherols, and plastoquinones (Fig. 2). It has become evident that both pathways are heavily regulated at multiple levels as was discussed in two recent reviews by Hemmerlin and coworkers [4, 5]. In addition to the transcriptional regulation of MVA and MEP pathway genes and their different paralogues, isoprenoid-pathway fluxes are controlled at posttranscriptional/-translational levels and by feedback regulation. Recent studies have given a more global view of the dynamics and networks of the core isoprenoid pathways and the regulation of metabolic flux during plant development and in response to external stimuli (reviewed in [6, 7]). Therefore, this chapter primarily gives an overview of both pathways with some emphasis on those in Arabidopsis and provides updates on the different modes of regulation.
Enzymatic steps of the MVA and MEP pathways and their regulation in isoprenoid precursor biosynthesis. Colored dots indicate the different levels of regulation for each enzyme according to the current status of knowledge [5]: green—transcriptional, purple—posttranscriptional, yellow—translational, red—posttranslational including feedback modulation. Other selected external and internal regulatory factors and posttranslational modifications of main regulatory enzymes are depicted as mentioned in the text. One or more gene paralogues as described from different plant species [4] are indicated by (P). Arrows indicate preferred trafficking of isoprenoid precursors between the cytosol and plastids in light (white) and dark (black) exposed tissues. Abbreviations for enzymes (red) and metabolites (black—MVA pathway; blue—MEP pathway) are as described in the text
Terpenoid biosynthetic pathways and their subcellular organization. Enzymes are marked in red; specialized terpenoids are marked in blue; all other intermediates and terpenoid end products are in black. Solid and dashed arrows indicate single and multiple enzymatic steps, respectively. Colored ovals indicate the homodimeric and heterodimeric composition of prenyltransferases involved in specialized terpenoid biosynthesis: GPS1—GGPS type, GPS2—heterodimer (heterotetramer) with large subunit (LSU) and small subunit (SSU I or SSU II), GPS3—homodimeric SSII type [136], GPS4—PPS type. Short-chain cis-prenyltranferases are marked in green. Abbreviations not mentioned in the text: IPS isoprene synthase; GRR geranylgeranyl reductase; OPS oligoprenyl diphosphate (OPP) synthase; SPS solanesyl diphosphate (SPS) synthase; SQS squalene synthase
2.1 MVA and MEP Pathways—A Brief Summary of Their Biosynthetic Steps
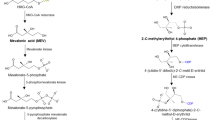
The MVA pathway in plants (Fig. 1a) consists of six steps and starts with the Claisen-type condensation of two molecules acetyl-CoA to acetoacetyl-CoA (AcAc-CoA) catalyzed by acetoacetyl-CoA thiolase (AACT). In a subsequent aldol condensation reaction catalyzed by HMG-CoA synthase (HMGS), AcAc-CoA is combined with a third molecule of acetyl-CoA to form the C6-compound S-3-hydroxy-3-methylglutaryl-CoA (HMG-CoA). Response to different stresses, feedback regulation, and the role of HMGS in sterol metabolism (see below) support a key function of HMGS in the MVA pathway. In the following rate-limiting step, HMG-CoA reductase (HMGR) catalyzes the conversion of S-HMG-CoA to R-mevalonate in two NADPH-dependent reduction steps. All plant HMGR proteins are membrane-bound with two membrane-spanning sequences and a highly conserved catalytic C-terminal domain. The presence of ER-specific retention motifs indicates a primary association of the membrane-spanning domain with the ER, whereas the N-terminal and C-terminal ends are positioned on the cytosolic side [8–13]. The association of HMGR to membranes seems to regulate its activity negatively, thereby limiting the accumulation of terpenoid end products such as sterols (e.g., [14, 15]). Many studies have reported on the critical regulatory role of HMGR in the biosynthesis of phytosterols, triterpenoids, and sesquiterpenoid phytoalexins, although flux control often involves additional downstream enzymes such as sesquiterpene synthases (e.g., [16–21]). MVA produced by HMGR is finally converted into IPP via three enzymatic steps: two ATP-dependent phosphorylation steps, catalyzed by mevalonate kinase (MK) and phosphomevalonate kinase (PMK), and an ATP-driven decarboxylative elimination catalyzed by mevalonate diphosphate decarboxylase (MVD or MPDC).
The MEP pathway (Fig. 1b), which occurs in all photosynthetic eukaryotes and in cyanobacteria, apicomplexan protozoa, and most eubacteria [22–25] consists of seven enzymatic steps. In the first reaction, 1-deoxy-d-xylulose 5-phosphate (DXP) is formed by DXP synthase (DXS) from (hydroxyethyl) thiamine diphosphate, which is derived from pyruvate, and glyceraldehyde-3-phosphate (GAP) in a transketolase-like condensation. Plant DXS enzymes carry a highly conserved thiamine phosphate binding domain and are divided in the class-I type enzymes with primary expression in photosynthetic and floral tissues and the class-II type enzymes with more distinct roles in specialized metabolism (see below). Numerous studies have confirmed that DXS functions as an important regulatory and rate-limiting enzyme in the biosynthesis of plastidial terpenes [26–31]. Consequently, DXS mutants such as those of the single functional Arabidopsis class-I type DXS gene (DXS1) exhibit albino phenotypes [32–34].
The enzyme 1-deoxy-d-xylulose 5-phosphate reductoisomerase (DXR) catalyzes the second step of the MEP pathway, in which DXP is converted into 2-C-methyl-d-erythritol 4-phosphate (MEP) by an intramolecular rearrangement of DXP into 2-C-methyl-d-erythrose 4-phosphate, followed by an NADPH-dependent reduction [35, 36]. The reaction can be specifically inhibited by fosmidomycin, a structure analogue of the DXR substrate [37–39] thereby blocking the biosynthesis of downstream plastidial terpene biosynthesis [40–42]. The reaction catalyzed by DXR is in some cases considered a rate-limiting step depending on the species, tissue, and developmental stage. In Arabidopsis, DXR1 is expressed in different plant organs [36] and dxr mutants show, similar to those of DXS1, an albino phenotype and deficiencies in gibberellin and abscisic acid (ABA) biosynthesis [43].
MEP is further converted in a CTP-dependent reaction to 4-diphosphocytidyl-2-C-methyl-d-erythritol (CDP-ME) by the enzyme 4-diphosphocytidyl-2-C-methyl-d-erythritol synthase (MCT or IpsD) [44, 45]. Phosphorylation of CDP-ME by the enzyme 4-diphosphocytidyl-2-C-methyl-d-erythritol kinase (CMK, IspE) then leads to the formation of 4-diphosphocytidyl-2-C-methyl-d-erythritol 2-phosphate (CDP-ME2P) [46–48], which is subsequently cyclized by 2-C-methyl-d-erythritol 2,4-cyclodiphosphate synthase (MDS, IspF) into 2-C-methyl-d-erythritol 2,4-cyclodiphosphate (MEcPP) upon loss of CMP. In the last two steps of the MEP pathway, the enzyme 4-hydroxy-3-methylbut-2-enyl diphosphate synthase (HDS, IspG) first converts MEcPP in a two-electron reduction to 4-hydroxy-3-methylbut-2-enyl diphosphate (HMBPP). In a final branching step, HMBPP is converted by 4-hydroxy-3-methylbut-2-enyl diphosphate reductase (HDR, IspH) to a mixture of IPP and DMAPP with a ratio of 5 to 6:1 [49–51].
Mutants of MCT, MDS, and CMK exhibit similar albino phenotypes and downregulation of photosynthetic genes [52, 53]. Likewise, hds and hdr-1 mutants have defects in chloroplast development [54, 55]. Interestingly, a partial loss-of-function mutant of Arabidopsis HDS, hds-3 (csb3), was shown to be more resistant to biotrophic pathogens suggesting a link between the MEP pathway and plant defense responses [56].
2.2 Differential Expression of MVA and MEP Pathway Isozymes
Several enzymes of the MVA and MEP pathways, especially those with important regulatory roles, are encoded by small gene families, which allow for functional redundancy and divergence (summarized in [4]; Fig. 1). In the MVA pathway, paralogues have been identified for AACT, HMGS, HMGR, and MPDC, whereas the MEP pathway enzymes DXS, DXR, MCT, CMK, MDS, or HDR were found to be encoded by two or more isogenes [4]. The different roles of many of the MVA and MEP pathway isozymes depend on their expression in specific cellular tissues and are often divided into essential functions to provide terpenoid precursors in primary metabolism, growth, and development, and more specific functions in stress response and specialized metabolism. For example, in Brassica juncea, HMGS is represented by a four-member gene family. Two genes are highly expressed at early stages of floral development [57–59] and play a role in reproduction, as was also shown for the single HMGS gene in Arabidopsis, [60] whereas expression of the two other paralogues is restricted to leaves [58, 59].
Notably, paralogues of the HMGR family exhibit different developmental and tissue-specific expression patterns and can be distinguished by their response to endogenous molecules such as phytohormones and sterol metabolites as well as external stimuli that include light, wounding, elicitor treatment, and pest and pathogen attack (Fig. 1; reviewed in [4]). The differential expression of HMGR isozymes, as demonstrated by early studies of the HMGR gene families in Solanaceous plants (tomato, potato) [61, 62], is important for channeling and counterbalancing carbon flux to the different downstream pathways of stress response or development. This functional differentiation, however, does not seem to occur in all plants inasmuch as both HMGR genes in Arabidopsis do not respond to stress but are essential for the production of sterols for cell elongation, senescence, gametophyte development, and fertility [63, 64].
Functional divergence of MEP pathway genes has been primarily observed in the DXS gene family. Class II-type DXS genes respond to biotic interaction and are induced in the biosynthesis of apocarotenoids upon mycorrhizal colonization in legumes and other plant families [30, 65, 66]. Several studies also demonstrated that type II DXS genes are induced in response to pathogen and herbivore attack in association with the production of specialized metabolites (summarized in [4]).
2.3 Metabolic Regulation and Networks
There is clear evidence for the role of pathway intermediates and downstream metabolites in the regulation of the core terpenoid biosynthetic steps at transcriptional and posttranslational levels (Fig. 1). Feedback inhibition by free CoA has been demonstrated for AACT and HMGR and for the enzymatic products of HMGS [59, 67, 68]. Furthermore, plant MKs respond to feedback inhibition by the prenyl diphosphates, IPP, DMAPP, geranyl diphosphate (GPP), and farnesyl diphosphate (FPP) that modulate enzyme activity by acting as competitive inhibitors of ATP [69]. Similarly, in vitro feedback inhibition was found for a DXS protein from poplar by IPP and DMAPP and a structural analysis suggested possible binding of the prenyl diphosphates to the enzyme in competition with its thiamine pyrophosphate substrate [70]. This feedback inhibition has also been supported in vivo by recent metabolic flux studies in poplar [71].
The complexity of the regulatory network also becomes apparent when metabolic disturbances and changes in metabolic flux generated by overexpression or reduced expression of genes of the core isoprenoid pathways promote pathway feedback or feedforward signals that modify the expression of up- or downstream genes. For example, overexpression of B. juncea wild-type and mutated HMGS1 in Arabidopsis caused an upregulation of HMGR and genes in sterol biosynthesis such as sterol methyltransferase 2, delta-24 sterol reductase, and C-22 sterol desaturase, which led to an elevated sterol content in leaves and seedlings and increased stress tolerance [57]. A similar response was observed for HMGS overexpression in tobacco resulting in improved sterol content, growth, pod size, and seed yield [72]. Conversely, knockdown of AACT2 expression led to lower levels and altered profiles of sterols and caused reduced expression of downstream genes encoding FPP synthases and sterol methyltransferase [73]. HMGR activity also exhibits a positive feedback response to downstream metabolic changes such as reduced cycloartenol levels in transgenic tobacco expressing sterol methyltransferase type 1 (SMT1) and the depletion of endogenous sterols due to the inhibition of squalene synthase [74, 75].
The simultaneous response of several genes to pathway perturbations is further observed in mutants of the MEP pathway. For instance, silencing of CMK in Arabidopsis causes upregulation of MCT, MDS, and HDS expression [76]. Moreover, in rice, MEP pathway genes were found to be coexpressed with downstream genes in carotenoid and phytyl biosynthesis [77]. In line with these observations, detailed transcriptional coexpression network analyses in Arabidopsis demonstrated that gene modules in both MVA and MEP pathways are coregulated together with genes of downstream pathways and these findings have set the stage to identify regulatory elements of these gene modules [78–80]. Consequently, cis elements were mapped showing that the promoters of the Arabidopsis genes DXS, DXR, CMK, HDR, and phytoene synthase share a cis-regulatory element interacting with RAP2.2, a member of the ethylene response factor B-2 subfamily [6].
In conjunction with their regulation by light (see below), MEP and MVA pathways respond to regulators in sugar metabolism. Arabidopsis mutants of pleiotropic regulatory locus 1 (PRL1), a global regulator of sugar, stress, and hormone responses, accumulate MEP pathway-derived end products (Fig. 1a) [81]. The same mutants have reduced HMGR activity but no change in HMGR transcript or protein because of posttranslational modification. PRL1 inhibits the SNF1 (sucrose nonfermenting)-related protein kinase 1 (SnRK1), which negatively regulates HMGR1 by phosphorylation and inactivation of the catalytic domain (Fig. 1a) [82]. HMGR1 is also negatively regulated during normal development and in response to salt stress by protein phosphatase 2A (PP2A), which dephosphorylates the HMGR protein (most likely at a site different from the phosphorylation by SnRK1; Fig. 1a) [83]. Modulation of HMGR transcripts at the initiation of translation [84] and glycosylation of HMGR isoforms [11], respectively, have been discussed previously as other mechanisms of posttranscriptional or posttranslational regulation of stress-induced HMGR genes.
There are several possible connections of the isoprenoid pathway to other metabolic routes by delivery and competition for carbon precursors (e.g., amino acid degradation) [4], which will require further attention to gain a more comprehensive understanding of flux in terpenoid biosynthesis. A link of isoprenoid metabolism with lipid biosynthesis was described by Nieto et al. [85], who found that inhibition of sphingolipid biosynthesis in Arabidopsis caused posttranslational downregulation of HMGR activity decoupled from HMGR transcript and protein levels and a reduction in sterol content. Recently, an unexpected simultaneous downregulation of flavonoid and terpenoid metabolite levels was observed in trichomes of tomato mutants of the flavonoid biosynthetic enzyme chalcone isomerase (CHI) [86]. These results have led to several hypotheses about the regulatory connections between both pathways. It is possible that changes in the levels of flavonoids by accumulation (upstream of CHI) or depletion (downstream of CHI) modify terpenoid biosynthetic gene expression or directly inhibit biosynthetic and regulatory proteins [87, 88]. Based on previous findings, there is also the possibility that CHI itself might interact with proteins involved in terpenoid production or its regulation [89]. Furthermore, it will be important to examine regulatory factors that coordinate the metabolic flux through both pathways [90].
2.4 Regulation by Light and External Stimuli
Thanks to recent efforts to identify MVA and MEP pathway gene expression patterns by transcriptome and hierarchical cluster analyses it was shown that the genes of both pathways have opposite expression patterns during light or dark (Fig. 1) [6]. Whereas exposure to light leads to the downregulation of MVA pathway genes and reduced levels of sterols [79], it stimulates transcript accumulation of MEP pathway genes and genes in the carotenoid and chlorophyll biosynthetic pathways such as PSY (phytoene synthase) and HEMA1 (glutamyl-tRNA reductase), which are essential for chloroplast differentiation [79, 80, 91–94]. Light also upregulates tocopherol, and plastoquinone biosynthetic genes such as VTE3 (vitamin E defective 3) [79]. The results are supported by studies that observed an increased carbon flux through the MEP pathway under enhanced light conditions by measuring the accumulation of MEcDP when 2-C-methyl-d-erythritol 2,4-cyclodiphosphate reductase activity was inhibited [95]. In contrast to the upregulation by light, expression of MEP pathway genes with the exception of HDR [55] is reduced during light–dark transition [6]. Dark exposure can induce HMGR activity as was shown in ginseng where HMGRs play a regulatory role in triterpene ginsenoside biosynthesis [96]. The light-dependent response of Arabidopsis MEP and MVA pathway genes is controlled by phytochrome B (PHYB) because phyB mutants have enhanced transcript levels and enzyme activity of HMGR but reduced levels of MEP pathway products [92]. Consequently, phytochrome interacting factors (PIFs) of the basic helix–loop–helix (bHLH) transcription factor family were identified as regulators that are involved in the light control of MEP and carotenoid biosynthetic pathway genes [93, 97]. Turnover of the MEP pathway enzymes DXS and DXR was also found to be correlated with the activity of Clp, a major plastid stromal protease (Fig. 1b) [98].
Downregulation of MEP pathway enzymes in the dark provides a dilemma for the biosynthesis of carotenoids and gibberellins required for the development of etiolated seedlings. Supported by observations from treatments with the MEP pathway inhibitor fosmidomycin, Rodriguez-Concepcion and coworkers suggested that during seedling germination in the dark, prenyl diphosphates derived from the MVA pathway are transported into etioplasts for gibberellin and carotenoid synthesis prior to the induction of MEP pathway enzymes upon illumination [92]. Given the responses of MEP and MVA pathway genes in light and dark, it is not surprising that the expression of several genes is under circadian control [7]. Coexpression analyses in Arabidopsis photosynthetic tissue connect several MEP pathway genes with core circadian oscillators (LHY, CCA1, PRR9) whereas only AACT2 of the MVA pathway follows the expression of circadian regulators peaking in the dark [6]. However, in roots, expression of several MVA pathway genes such as HMGR1 is correlated with that of circadian regulators (TOC1, TIC) showing clear differences in the circadian control of early pathway genes in above- and belowground tissues. Interestingly, in triple mutants of the TOC1 related pseudo-response regulator (PRR) proteins PRR9, PRR7, and PRR5, genes and metabolites of carotenoid, chlorophyll, and tocopherol pathways are upregulated, which suggests a function of these proteins as negative regulators of the MEP pathway-dependent metabolic routes [99]. To what extent the oscillation of MVA and MEP pathway gene transcripts directly corresponds to changes in enzyme activity and downstream metabolites requires further attention. In snapdragon flowers, the rhythmic emission of volatile monoterpenes in plastids and sesquiterpenes in the cytosol depends on the MEP pathway that is controlled by the circadian clock [100].
In addition to their differential response to light, MVA and MEP pathways respond to multiple other external stimuli at gene transcript and posttranslational levels (Fig. 1; summarized in [4]). To support the production of terpenoids for protection against temperature stress, carbon flux through the MEP pathway increases under elevated temperatures [95]. In the MVA pathway, not only HMGR but other enzymes such as AACT show induced responses under abiotic stress and appear to be involved in MVA pathway-mediated abiotic stress adaptation [68]. Changes in redox state also directly affect MVA and MEP pathway enzymes. Both HDS and HDR, which function as iron–sulfur reductases, have been identified as targets of the redox protein thioredoxin [101, 102] and thioredoxin-dependent regulation has also been suggested for DXR [102]. Moreover, it has been shown that HDS can receive electrons directly through the photosynthetic electron-transport chain via ferredoxin without any reducing cofactor, which is different from the flavodoxin/flavodoxin reductase and NADPH-dependent reducing system of HDS in bacteria [103].
Biotic stress such as pathogen attack often upregulates individual genes of HMGR families to direct flux toward the production of sesquiterpene phytoalexins under simultaneous downregulation of squalene synthase and sterol biosynthesis [62, 104]. Studies in tobacco showed that the regulation of pathogen-activated expression of HMGR involves the MEK2-SIPK/WIPK MAP kinase cascade [105, 106]. Another example highlights the importance of HMGR in root nodule development. The HMGR1 protein of Medicago truncatula directly interacts with NORK, which is a receptor-like kinase required for Nod factor signaling. Reduced expression of HMGR1 in transgenic plants causes a severe decrease of root nodulation [107].
2.5 Regulation and Metabolite Exchange Across Subcellular Compartments
The compartmentalization of MEP and MVA pathways and associated downstream pathways allows for the subcellular regulation and coordination of photosynthesis-dependent and independent terpenoid biosynthetic routes. Despite the general notion that the MVA pathway enzymes are located in the cytosol or associated with the ER, peroxisomes have been discussed as localization sites for AACT (particularly AACT1 in Arabidopsis), PMK, and MVD based on the prediction of peroxisomal PTS targeting peptides and transient protein peroxisome import studies in Catharanthus roseus cells [108–110]. For MVD1 in Arabidopsis, however, mass spectrometry analysis suggests a cytosolic localization and MVD2 is predicted to reside in the cytosol [6]. In the absence of additional evidence for a partial localization of the MVA pathway in peroxisomes and possible transporters of isoprenoid precursors between the compartments, our current view on the subcellular organization of the MVA pathway remains incomplete.
The exchange of intermediates between the cytosol and plastids is usually not sufficient to rescue Arabidopsis mutants of biosynthetic enzymes in the MVA or MEP pathways [31, 43, 64]. However, studies on dxs2 mutants in tomato suggested that both pathways can, to some extent, compensate each other [66]. Moreover, in numerous cases, some degree of exchange of isoprenoid intermediates between plastids and the cytosol has been demonstrated based on the application of MEP and MVA pathway-specific inhibitors and the incorporation of stable-isotope precursors in primary and specialized terpenoid metabolites (e.g., [100, 111–117]. There is frequent evidence for trafficking of isoprenoid intermediates from the plastid to the cytosol in photosynthetic tissues (e.g., [113]). However, the contribution of the MVA pathway to the biosynthesis of plastidial isoprenoids can be substantial in the absence of light as was demonstrated by Opitz et al. [118] in roots of cotton seedlings or in dark-grown Arabidopsis seedlings [92].
To date, no specific transporters of isoprenoid precursors have been identified in the plastid membrane. The export of IPP from plastids to the cytosol was suggested to proceed by a plastidial proton symport system [119]. Studies by Flügge and Gao [120] indicated that IPP is not transported by plastidic phosphate translocators but depends on phosphorylated counter-substrates. In addition to the transport of IPP, there is evidence that longer prenyl diphosphates such as GPP and FPP are moved from plastids to the cytosol in tomato [121], the grape berry exocarp [122], and glandular trichomes of Stevia rebaudiana [116]. Genomic and proteomic analyses of single cells such as trichomes could be a promising approach to identify the isoprenoid transporter machinery between both compartments.
Despite some degree of exchange of isoprenoid intermediates between the plastid and the cytosol, the spatial separation of terpenoid biosynthetic pathways has been of benefit for the engineering of terpenoid end products. Expression and targeting of an FPP synthase and sesquiterpene synthase to plastids in tobacco did prevent carbon flux competition with sterol biosynthesis in the cytosol and promoted sesquiterpenoid yields by a thousandfold [123]. The same approach was successfully applied to produce high levels of the triterpene squalene in plastids and in tobacco trichomes although the latter case came at the cost of severely reduced growth [124]. Efforts have also been made to insert the entire MVA pathway in the tobacco chloroplast genome resulting in increased levels of mevalonate and carotenoids, but also squalene and sterols [125].
As mentioned above, expression of the MEP and MVA pathway genes is coordinately regulated by external stimuli. Other interdependent mechanisms of regulation between the pathways have been detected at posttranslational levels. Recent studies in tobacco demonstrated that blocking MEP pathway-dependent protein geranylgeranylation by treatment with the monoterpene S-carvone suppresses signaling to induce the MVA pathway-dependent formation of the sesquiterpene phytoalexin capsidiol [126]. Other possible roles of multicompartment networks in regulating the MVA pathway have been addressed by Verbitskiy et al. [127]. Work by these authors on proteins involved in RNA editing suggests that retrograde signaling between mitochondria and the cytosol might modify MVA pathway activity and, according to Tang et al. [128], this interaction seems to involve the mitochondrial respiratory pathway. Most notably, the MEP pathway intermediate, MEcPP, was found to function as a retrograde signaling molecule between plastids and the nucleus. MEcPP elicits the expression of stress-responsive nuclear-encoded plastidial proteins which suggests that the MEP pathway functions in stress sensing and coordinating stress-induced nuclear genes [129].
3 Isomerization and Condensation of the C5 Building Blocks
The construction of terpenoids with more than five carbons requires a sufficient supply of IPP and its more reactive, electrophilic isomer DMAPP. Therefore, IPP derived from the MVA pathway needs to be converted to DMAPP by the activity of an IPP isomerase (IDI; Figs. 1 and 2). Type I IPP isomerase isoenzymes in plants have been localized to mitochondria and plastids and shorter isoforms have been predicted to remain in the cytosol [130]. In analogy to mammalian cells, an alternative localization of IPP isomerases in peroxisomes has been discussed [108] but additional evidence for the role of peroxisomes in plant isoprenoid metabolism is needed. Although the formation of DMAPP from IPP derived from the MVA pathway is essential for downstream reactions in the cytosol and mitochondria, IPP isomerization seems less important in plastids where both C5 building blocks are produced by the MEP pathway. However, plastidial IPP isomerase activity might be necessary to produce an optimal ratio of IPP and DMAPP for the downstream condensation reactions and to provide precursors for a possible transport to the cytosol.
In the second major stage of terpenoid biosynthesis, IPP and DMAPP units are fused by the catalytic activity of prenyltransferases (isoprenyl diphosphate synthases) to form prenyl diphosphates as the linear central precursors of all terpenoids (Fig. 2). The initial reaction catalyzed by a prenyltransferase is a head-to tail (1′–4) condensation of IPP with the allylic cosubstrate DMAPP based on an ionization–condensation–elimination mechanism to produce a C10-allylic diphosphate. Additional rounds of head-to-tail condensation of the allylic product with more IPP units lead to the formation of short-chain (C15–C25), medium-chain (C30–C35), and long-chain (C40–Cn) prenyl diphosphates. The cis- or trans-stereochemistry of the double bonds of the prenyl diphosphate product determines whether the enzyme operates as cis-prenyltransferase or trans-prenyltransferase, which belong to families of structurally unrelated enzymes [131]. Much knowledge has been gained on the biochemistry and evolution of short-chain trans-prenyltransferases, which synthesize C10-geranyl diphosphate (GPP), C15-trans,trans-farnesyl diphosphate ((E,E)-FPP), or C20-all-trans-geranylgeranyl diphosphate (all-trans-GGPP) as the main precursors in terpenoid metabolism, although more recent work has discovered similar roles of previously undetected short-chain cis-prenyltransferases (see below).
3.1 Geranyl Diphosphate Synthases
As a precursor in the biosynthesis of C10-monoterpenoids, GPP is synthesized from IPP and DMAPP by the activity of GPP synthase enzymes (GPSs), which are usually targeted to plastids (Fig. 2). Different classes of homodimeric and heterodi/tetrameric GPSs have been identified in plants [132–136] (Fig. 2). A heterotetrameric GPS from peppermint was the first GPS to be discovered in plants [137] and since then related heterodimeric proteins have been found in a variety of other species such as Anthirrinum majus, Clarkia breweri, and Humulus lupulus [137–139]. The enzymes consist of a large subunit (LSU), which has significant homology (~50 %) to GGPP synthases (GGPS, see below) and can exhibit GGPP synthase activity as a recombinant protein, and a small subunit (SSU I) that shares only ~20 % sequence similarity with homomeric prenyltransferases and is functionally inactive. It is generally thought that binding of SSU I modifies the activity of the LSU to produce GPP. The importance of the physical interaction of both subunits to make GPP has been confirmed by structural analysis of the heterotetrameric GPS from peppermint [134]. In Arabidopsis, Wang and Dixon [139] identified a separate lineage of SSU (SSU II) genes encoding GGPS-related proteins (GGR). Arabidopsis GGR modifies the in vitro activity of GGPS 11 to produce GPP and contains two conserved CxxxC motifs that are essential for the interaction of both subunits [139]. In contrast to the role of SSU I-containing GPSs in monoterpene formation in peppermint or hops, the function of heterodimeric GPSs carrying SSU II subunits is less clear because of the absence of a tight correlation between protein expression and the biosynthesis of monoterpenes in different tissues [139].
Engineering of GPS activity has been achieved by the expression of GPS.SSU I from snapdragon in tobacco and tomato fruits. The expressed subunit recruits plastidial GGPS proteins to form functionally active heterodimeric GPS proteins [121, 140]. The study on tomato also revealed that GPP produced in plastids is exported to the cytosol, where it can be used for monoterpene biosynthesis [121]. However, the exchange of GPP between both compartments might be limited in the absence of engineered GPP pools as was shown for a bifunctional Arabidopsis monoterpene/sesquiterpene synthase (TPS02), which is located in the cytosol and produces sesquiterpenes but no monoterpenes in planta [42].
Homodimeric GPS enzymes have been described from angiosperms and gymnosperms [135, 136, 141, 142]. These proteins belong to different lineages and are evolutionarily related to GGPSs (see below). The existence of a homodimeric GPS in Arabidopsis has been discussed controversially. A single GPS1 gene was originally identified to encode a functionally active GPS enzyme [143]; however, more recently the GPS1 protein has been characterized as a multiproduct medium-/long-chain prenyl diphosphate synthase. The latter activity was observed when IPP was supplied in excess to the allylic substrates DMAPP, GPP, and FPP and was supported by the structural analysis of an active-site cavity with sufficient size to accommodate the medium-/long-chain products [144]. The GPS1 protein (renamed by Hsieh et al. as polyprenyl di(pyro)phosphate synthase, PPS) is targeted to plastids [143] where IPP and DMAPP are produced at ratios of approximately 5:1 by the MEP pathway. Thus, it is possible that this enzyme exhibits a PPS activity in vivo.
3.2 Farnesyl Diphosphate Synthases
Trans-FPP synthases (FPSs) catalyze the formation of (E,E)-FPP as a central precursor in the biosynthesis of terpene primary metabolites (phytosterols, brassinosteroids, dolichols, ubiquinones), for protein prenylation, and in the production of specialized metabolites such as sesquiterpenoids and triterpenoids (Fig. 2). As type I (eukaryotic) FPSs, plant trans-FPSs build a superfamily of homodimeric enzymes that are often encoded by small species-specific gene families (e.g., [145–147]). FPS isozymes of different size that are produced as a result of differential gene transcription have been localized to the cytosol or the mitochondria where they produce FPP pools for the biosynthesis of cytosolic and mitochondrial downstream products [148] (Fig. 2). Targeting of FPSs to peroxisomes has been discussed based on YFP fusion experiments in Catharantus roseus cells [149]. However, no peroxisomal targeting has been demonstrated for fluorescent FPS fusion proteins in Arabidopsis, which is consistent with results from proteomic studies of the cytosol and purified peroxisomes [150, 151].
As with the isozymes of the MEP and MVA pathways, it has been a primary interest to elucidate the possible functional differences of prenyltransferase isoforms. In Arabidopsis, the two FPS paralogues, FPS1 and FPS2, have overlapping expression patterns and can rescue each other’s loss, whereas double mutants are impaired in male genetic transmission and arrested at early embryo development [152]. However, there is no complete functional redundancy between the two isozymes inasmuch as FPS2 is the predominantly expressed isozyme in mature seeds and early seedling development, and FPS1 appears to be only expressed in the maternal seed coat [153]. Consequently, seeds of fps2 mutants have a reduced sterol content [152]. Keim et al. propose that the specific expression of FPS2 in mature seeds is related to its higher enzymatic activity and thermal stability. The authors further speculate that during early development of the embryo (in the absence of FPS2 expression), FPP might be imported from the seed tissue where FPS1 is expressed [153].
3.3 Geranylgeranyl Diphosphate Synthases
Similar to (E,E)-FPP, all-trans-GGPP synthesized by all-trans-GGPSs is a major branching point for several downstream terpenoid pathways in primary and specialized metabolism. These include the biosynthesis of carotenoids and their breakdown products (abscisic acid, strigolactones), chlorophylls, tocopherols, gibberellins, plastoquinones, and diterpenoids (all synthesized in plastids), geranylgeranylated proteins and poly-/oligoprenols (synthesized in the cytosol), and poly-/oligoprenols synthesized in the plastids and mitochondria (Fig. 2). Compared to FPSs, GPPS isozymes are represented by larger gene families. For example, the Arabidopsis genome contains 12 GGPS paralogues, of which 10 have been identified to encode functional GGPS proteins of most likely homodimeric architecture and with GGPP as the primary or sole product [154]. The different GGPS isozymes are located in the plastids, mitochondria, and the ER consistent with the subcellular compartmentalization of the diverse GGPP-dependent terpenoid pathways. With the exception of two of the Arabidopsis isozymes (GGPS1-mitochondrial, GGPS11-plastidial), which are expressed in the whole plant, the remaining family members exhibit distinct spatiotemporal expression patterns [154]. Seedling-lethal albino and embryo-lethal phenotypes are found in ggps1 mutants, indicating that GGPS1 has essential functions in development and the chlorophyll biosynthetic pathway [155]. Although possible redundant or more specific functions of most of the GGPS isozymes are not well understood, it is apparent that the divergence in the Arabidopsis GGPS gene family is the result of functional specialization and fine-tuning of metabolic pathways in different cellular compartments and in tissues at different developmental stages or under different environmental conditions.
Both FPPS and GGPPS proteins have been expressed in modules with sesquiterpene synthases and diterpene synthases, respectively, to engineer the biosynthesis of sesquiterpenoids and diterpenoids in microbial systems and in planta [123, 156]. Specifically, the buildup of FPP pools in plastids improved the precursor supply and allowed for a substantial increase in yield of the desired sesquiterpene products [123]. Other strategies to improve pathway productivity include generating combinatorial mutations in prenyldiphosphate synthase and downstream terpene synthases. For example, prokaryotic expression of pathway variants of a GGPPS and a terpene synthase, which produces a levopimaradiene diterpene precursor in ginkgolide biosynthesis, led to a more than 2,000-fold increase in the levels of the levopimaradiene product thereby stressing the importance of protein engineering in these approaches [157].
3.4 Chain Length Regulation and Evolution of Prenyltransferases
Structural analysis combined with random or site-directed mutagenesis has provided substantial insight to the chain length regulation of short-chain prenyltransferase products [158]. Based on crystal structures of several homodimeric FPPs and GGPS from eukaryotes and prokaryotes [159–165], short-chain prenyltransferases share a common protein fold composed of 13 α-helices with 10 helices surrounding the active site cavity. IPP and the allylic substrate are bound by two highly conserved aspartate-rich regions, a first DDx2-4D motif (FARM) and a second DDxxD motif (SARM), which are positioned on opposite walls of the cavity. Product chain length is in part regulated by amino acid residues upstream of the FARM motif (position –4, –5), which change the size of the hydrophobic substrate binding or elongation pocket of the polyisoprenoid chain [166, 167]. According to this mechanism, type I FPSs such as Arabidopsis FPS1 and FPS2 have a smaller binding pocket because of the presence of “bulkier” aromatic amino acid residues. In type II GGPSs, which comprise eubacterial and plant GGPSs, these aromatic amino acids are replaced by smaller residues such as alanine, serine, and methionine allowing the formation of a longer C20 chain. Studies of yeast GGPS indicated that chain termination at C20 depends on residues located deeper in the catalytic cavity [162]. Poulter and colleagues recently employed a large-scale bioinformatics approach combined with experimental enzyme characterization, protein crystallization, and computational modeling to predict the chain length specificity of a large number of putative polyprenyl transferases [168]. The approach, which resulted in a high rate of correctly predicted functions, largely supported the notion that steric hindrance in the elongation cavity is the main criterion determining chain length specificity. It is important to note that the study also suggested a chain-length–determining effect of “second shell” residues that are positioned in the vicinity of the residues lining the elongation pocket. Depending on their size, these neighboring residues may or may not provide flexibility for bulkier aromatic residues that protrude into the cavity to be moved or displaced by the growing polyprenyl chain [168].
Phylogenetic analyses of prokaryotic and eukaryotic prenyltransferases place plant FPSs in a clade with other eukaryotic FPSs that is distinct from a cluster containing plant GPS and GGPS proteins [158]. A comprehensive phylogenetic study of GGPS and GPS homologues of land plants and green algae demonstrated a lineage and species-specific expansion of GGPS families indicating gene duplication events and functional divergence [169]. The phylogeny shows several evolutionary transitions from proteins with GGPS to GPS activity. For example, gymnosperm homodimeric GGPSs, which form a distinct clade among plant GGPSs, can produce shorter prenyldiphosphates or synthesize exclusively GPP [133, 142]. In comparison to GGPSs from green algae and mosses that possess the FARM and SARM motifs and a conserved CxxxC motif, the gymnosperm GGPSs have acquired a second CxxxS (bifunctional GGPS) or CxxxC (GPS) motif. The two CxxxC motifs are characteristic of most proteins that are associated with GPS activity. Thus, they are present in the SSU I and SSU II subunits of heteromeric GPS proteins and critical in binding the LSU. The binding of both subunits limits access to the elongation cavity and terminates chain elongation at the formation of a C10-product [134]. SSU I and SSU II proteins have lost both aspartate-rich motifs or carry a mutated SARM, respectively, which is associated with the loss of prenyl diphosphate activity [135, 169]. Interestingly, an earlier study reported a flower-specific GPS from orchids with similarity to SSU II [136]. This protein lacks the SARM but maintains GPS activity as a homodimeric enzyme (Fig. 2).
Several proteins with homology to Arabidopsis PPS (former GPS1) have been reported from other plants and designated as homodimeric GPSs (Fig. 2). These proteins do not carry the CxxxC motifs and it remains to be determined whether they function as true GPSs in vivo or may exhibit medium-chain or long-chain polyprenyl diphosphate activity as was shown for the Arabidopsis enzyme. For instance, GPS activity was demonstrated for a protein in tomato, but assays were performed at a low IPP/DMAPP ratio [141]. Furthermore, silencing or mutation of this enzyme and of PPS in Arabidopsis resulted in dwarfed or embryo lethal phenotypes, which could be related to promiscuous GGPS activity to produce GGPP for gibberellin biosynthesis or the synthesis of longer precursors in plastoquinone biosynthesis. The formation of longer chain products by Arabidopsis PPS is also supported by the absence of aromatic amino acids near the FARM. Computational predictions such as those presented by Wallrapp et al. [168] should facilitate determining the chain length specificity of PPS homologues. In summary, GPS activity appears to be the result of promiscuity and neofunctionalization of GGPS (or PPS?) proteins in conjunction with the evolutionary adaptation of individual plant lineages to produce monoterpenes as constituents of floral scent or for chemical defense.
3.5 Cis-Isoprenyl Diphosphate Synthases
One of the surprising findings in the field of terpene biosynthesis in the past five years was the identification of short-chain cis-prenyltransferases (CPTs) and the conversion of their cis-prenyl diphosphate products to terpenoids by the activity of terpene synthases (see below). Prior to this discovery, it was generally believed that CPTs synthesize prenyl diphosphate products with a chain length of more than 50 carbons by using all-trans short-chain prenyl diphosphates as allylic primer substrates [170]. Such prenyltransferases in plants include enzymes that produce C70–C120 dehydrodolichol diphosphates or natural rubber (>C10,000) from (E,E)-FPP by head-to-tail condensations in a cis orientation [170–172]. Functional genomics studies of terpene biosynthetic genes in glandular trichomes of wild tomato then revealed the presence of a short-chain (Z,Z)-FPP synthase that produces (Z,Z)-FPP [173] (Figs. 2 and 3). Characterization of a nine-member CPT family in cultivated tomato gave additional evidence for short-chain enzyme activity by the identification of three genes encoding a neryldiphosphate (NPP) synthase (NDPS1 or SlCPT1, expressed in trichomes), a (Z,Z)-FPP synthase (SlCPT6, expressed in root and fruit), and a nerylneryl diphosphate (NNPP) synthase (NNDPS or SlCPT2, expressed in the stem), respectively [174] (Figs. 2 and 3). All three proteins are targeted to plastids [174]. Notably, the Z,Z-FPP pool produced by (Z,Z)-FPP synthase in trichome-specific plastids in wild tomato is used by plastidic sesquiterpene terpene synthases (santalene/bergamotene sesquiterpene synthase [173] and 7-epizingiberene synthase [175, 176]), which are related to diterpene synthases. Engineering of (Z,Z)-FPP synthase and 7-epizingiberene synthase in trichomes of cultivated tomato led to the production of 7-epizingiberene and increased resistance to herbivores [175]. NPP has been shown to be converted by a monoterpene synthase to β-phellandrene among other monoterpenes [177]. Consequently, coexpression of the NDPS1 enzyme with phellandrene synthase 1 was used successfully for metabolic engineering of monoterpene formation in tomato fruits [178]. Intriguingly, expression of NDPS1 alone led to the reduction of carotenoid levels in fruits because of feedback inhibition of GGPS by NPP. Based on these findings, it is plausible that NPP production is restricted primarily to trichomes to avoid inhibitory effects on carotenoid biosynthesis.
Structural diversity of terpenoid specialized metabolites and their precursors. a, b Examples of monoterpenoids and sesquiterpenoids produced by different Arabidopsis terpene synthases (AtTPS). c Structures of prenyl diphosphates produced by short-chain CPTs; NPP neryl diphosphate; NNPP, nerylneryl diphosphate (a, b from Tholl and Lee 2011) [217], thearabidopsisbook.org, Copyright American Society of Plant Biologists
The association of the tomato CPT genes with terpenoid biosynthetic gene clusters [174] clearly indicates adaptive functional specialization in the tomato CPT gene family to provide short-chain prenyl diphosphates for different terpene biosynthetic pathways including trichome-specific terpene biosynthesis. In line with these findings, a cis-type prenyltransferase was identified in lavender that catalyzes the head-to-middle condensation of two DMAPP molecules to synthesize lavandulol diphosphate, the precursor of lavendulol [179]. Furthermore, in the nine-member CPT gene family of Arabidopsis a multiproduct prenyltransferase (AtCPT6) has been identified that makes polyisoprenoid diphosphates with six to eight isoprene units as precursors of polyisoprenoid alcohols in roots [180].
As with trans-prenyltransferases, efforts have been made to determine amino acid residues that control the chain length specificity of CPTs [170]. Sequences of CPTs share five conserved regions and employ residues for substrate binding and catalytic activity that are different from those of trans-prenyltransferases [170]. Kang et al. [181] exploited accession-specific sequence differences of NDPS and (Z,Z)-FPP synthase in tomato coupled with homology modeling and site-directed mutagenesis to identify four residues in region II that are important for product specificity. These residues are part of helix II, which, together with helix III, lines a hydrophobic cleft that influences product chain length [182, 183].
4 Conversion of Prenyl Diphosphates and Terpene Synthase Function and Regulation
Trans- and cis-prenyldiphosphates are the entry points to various downstream primary and specialized terpenoid biosynthetic routes in plastids, mitochondria, and the cytosol (summarized in Fig. 2). It is beyond the scope of this chapter to address all of these pathways and the reader is referred to other chapters in this series (e.g., carotenoid biosynthesis) or more specialized recent reviews in the field.
The tremendous diversity of terpenoids in specialized metabolism can to a large extent be attributed to the activity of terpene synthases (TPSs; Fig. 3a). TPS enzymes have, therefore, become a focus point of in planta and heterologous metabolic engineering of terpenoid end products with use as pharmaceuticals, flavors, biofuels, or plant chemical defenses [184] (see other chapters in this series). The TPS superfamily, which is divided into eight subfamilies (TPSa–h), comprises a large and still growing number of enzymes from almost all taxa in the plant kingdom [185]. TPSs convert acyclic C5 to C20 cis- or trans-prenyl diphosphate intermediates into C5-hemiterpenes such as isoprene, C10-monoterpenoids, C15-sesquiterpenoids, or C20-diterpenoids (Fig. 2). The primary enzymatic products are in most cases acyclic or cyclic hydrocarbons (Fig. 3a) that are frequently modified by secondary enzymatic reactions such as hydroxylation, peroxidation, methylation, acylation, glycosylation, or cleavage to produce biologically active end products of even larger structural diversity [186]. TPS enzymes facilitate adaptations of terpene metabolism to the changing environment because their promiscuous activity often results in the production of more than a single compound (e.g., [187]) and TPS proteins easily acquire new catalytic properties by minor structural changes [187–192].
Mechanistically, TPS proteins are divided into class I and class II enzymes. The enzymatic reaction catalyzed by class I TPSs starts with the ionization of the prenyl diphosphate substrate by a divalent cation-dependent subtraction of the diphosphate moiety. The produced carbocation intermediate then enters different reactions that can include cyclizations, hydride shifts, and rearrangements prior to a termination of the reaction by proton loss or the addition of a nucleophile such as water [193] (Fig. 3). By contrast, class II TPSs, which include oxidosqualene cyclases (see below) and diterpene synthases, catalyze the ionization of their substrate by adding a proton to an epoxide ring or via protonation at the 14,15-double bond of GGPP, respectively. Class II diterpene synthases that fall into this category are ent-copalyl diphosphate (CPP) synthases (CPSs), which are involved in gibberellin and phytoalexin biosynthesis [194] (Fig. 2). In the gibberellin biosynthetic pathway, CPSs catalyze a protonation-induced bicyclization of the substrate GGPP to form ent-CPP, which is further ionized and converted to ent-kaur-16-ene by a class I ent-kaurene synthase (KS) activity. Detailed genomic studies of land plants revealed that the gibberellin biosynthetic pathway gave rise to the biosynthesis of an array of specialized labdane-related diterpenoids largely by gene duplication and divergence of CPS and KS homologues [194]. The ability to produce kaurene arose early in land plant evolution as can be assumed from the identification of a bifunctional classII/I CPS/KS in the moss Physcomitrella patens, which catalyzes the formation of ent-kaurene (and 16-hydroxykaurene) via a CPP intermediate in the biosynthesis of kaurenoic acid [195, 196]. Similar class II/I diterpene synthases such as abietadiene synthase occur in gymnosperms and can be considered early diterpene synthases. These enzymes produce (+)-CPP from GGPP prior to an ionization-initiated cyclization of (+)-CPP to the diterpene product [197]. An interesting new view on the evolution of plant TPS genes comes from a genomic study of a large TPS gene family in the fern Selaginella moellendorffii [198]. Two distinct types of TPS genes were identified: a group of diterpene synthases that represent a new plant TPS-h subfamily, and, surprisingly, a group of monoterpene synthases and sesquiterpene synthases that are more closely related to microbial TPSs and may be the first indication for a horizontal gene transfer of TPS genes [198].
It should be noted here that, recently, a new mechanism for the enzymatic formation of cyclic terpenes was discovered in the iridoid monoterpene biosynthetic pathway [199]. Iridoids have pharmaceutical and antibacterial activities and are also produced by aphids as pheromones [200, 201]. The iridoid synthase from Catharantus roseus is a short-chain reductase that most likely generates a C5-iridoid ring in the linear monoterpene 10-oxogeranial substrate by coupling a reduction step with a cyclization step via a Diels–Alder cycloaddition or a Michael addition [199]. This exciting finding may open the way for future discovery of similar reductase-type terpene cyclases in plants and other organisms.
More insight to the evolution of “regular” TPS enzymes has been gained from the analysis of an increasing number of crystal structures including those from an isoprene synthase [202], monoterpene synthases [203–206], sesquiterpene synthases [207, 208], a class I diterpene synthase (taxadiene synthase [209]), a class II CPP synthase [210, 211], and a class II/I diterpene synthase (abietadiene synthase [212]). Comparisons of the assembly of a class I type α-domain and class II type β and γ domains led to the prediction of an evolutionary scenario according to which an ancestral bifunctional classII/classI diterpene synthase (consisting of all three domains with a functional α- and β-domain) similar to the CPS/KS enzyme of P. patens gave rise to class II type diterpene synthases (consisting of all three domains with a functionally active β-domain and an inactive α-domain) and class I type TPSs (consisting of a nonfunctional β-domain and a functionally active α-domain) [213, 214]. A functionally active class I α-domain carries the highly conserved aspartate-rich motif, DDxxD, and a less conserved NSE/DTE motif, which are located on opposite sides of the entrance of the catalytic side and help position the diphosphate substrate by binding of a trinuclear magnesium cluster [215]. By contrast, functional class II β-domains carry a conserved DxDD motif, which is required for protonation-initiated carbocation formation [213].
Although TPS enzymes may convert more than one prenyl diphosphate substrate in vitro, their function in vivo is largely determined by the substrate pool that is available in the respective cellular compartment. In this regard, TPS enzymes localized in plastids generally produce monoterpenoids or diterpenoids from plastidial GPP and all-trans-GGPP, respectively, whereas TPSs in the cytosol primarily convert (E,E)-FPP to sesquiterpenes (or squalene in the biosynthesis of C30 terpenes). However, this general rule has recently been challenged by the discovery of plastidial (Z,Z)-FPSs and sesquiterpene synthases in tomato, the latter of which are more closely related to kaurene synthases in the TPS-e subfamily [177].
The existence of medium-size to large TPS families in Arabidopsis and many other plant species strongly supports the notion that TPS genes evolve by gene duplication and neofunctionalization [185, 216, 217]. Such duplication events combined with relocation in the genome can include other genes that encode modifying enzymes such as cytochrome P450s, and thus lead to the assembly of gene clusters. From the first discovery of a thalianol triterpene biosynthetic gene cluster in Arabidopsis [218], several such clusters have been found in the arabidiol, marneral, and avenacin triterpene biosynthetic pathways in Arabidopsis and oat, respectively [219, 220] (Sohrabi et al. in preparation), and for the biosynthesis of labdane-related diterpenoids in rice [221] or monoterpenoids and sesquiterpenoids in tomato [222]. The triterpene biosynthetic clusters carry genes for oxidosqualene cyclases (OSCs), which catalyze the cyclization of oxidosqualene to one or more cyclic triterpene alcohols via formation of a carbocationic intermediate [223, 224]. Coexpression with other cluster genes (e.g., P450s, desaturase, acyltransferase) in an operon-like manner then allows a consecutive derivatization of the triterpene precursor [218–220]. The evolutionary forces driving this coordinated gene cluster assembly are believed to be twofold. Clustering of genes for pathway building facilitates the regulation of multiple genes at the level of chromatin and/or prevents the accumulation of possible cytotoxic products [219, 225, 226]. However, a strict coregulation of gene expression does not seem to be the case in all clusters as was shown for a diterpene biosynthetic cluster in rice containing P450s that are differentially regulated and function in two different pathways [194].
Clusters that exhibit a coordinated expression of their genes have allowed the identification of putative key regulators such as in the case of the basic leucine zipper transcription factor, OsTGAP1, which is involved in regulating a diterpenoid biosynthetic gene cluster in rice [227]. Another transcription factor that was identified previously to regulate terpene biosynthetic genes positively is a WRKY transcription factor in cotton, GaWRKY1, which regulates the transcription of a sesquiterpene synthase gene in the gossypol biosynthesis pathway [228]. More recent studies on Artemisia annua suggest that APETALA2/ethylene-response factors (AP2/ERF) are positive regulators of biosynthetic genes in the formation of the sesquiterpene artemisinin, an insect deterrent and antimalaria drug produced in leaf glandular trichomes [229]. However, these studies thus far do not place the identified transcription factors into regulatory networks related to development and cell specification.
A better understanding of the regulatory networks controlling terpene volatile formation has been gained in the process of flower maturation in Arabidopsis. Two R2R3 MYB transcription factors, MYB21 and MYB24, were identified that promote gynoecium growth and nectary development and positively affect expression of the major floral (E)-β-caryophyllene sesquiterpene synthase TPS21 [230]. Both MYB TFs respond positively to jasmonic acid (JA), the levels of which are induced by the auxin response factor 6 (ARF6) and ARF8, both master regulators of flower maturation. TPS21 and the second floral sesquiterpene synthase, TPS11 [187], also respond more directly to JA by the direct binding of their promoters to the bHLH transcription factor MYC2 [231], which is a central regulator of the JA signaling pathway in developmental and stress responses [232, 233]. In addition, TPS21 and TPS11 gene expression is indirectly regulated by gibberellins through the binding of DELLA proteins (gibberellin signaling repressors) [231].
Similar to the tissue-specificity of terpene formation in flowers, terpene-specialized metabolism in roots appears to be a highly coordinated cell type-specific process. Genes of the thalianol and marneral triterpene biosynthetic gene clusters are coexpressed primarily in the root epidermis [218, 219]. Likewise, 14 genes of the Arabidopsis TPS family are expressed in different root tissues. For example, a recently identified rhizathalene diterpene synthase (TPS08; Fig. 4) was found to be primarily expressed in the root stele (see below) [234]. In addition, two 1,8-cineole monoterpene synthase genes are constitutively expressed in the stele of the root elongation zone and differentiation/maturation zone and in the epidermis and cortex of more mature roots; a similar expression pattern has been observed for two closely related (Z)-γ-bisabolene sesquiterpene synthases [188, 235]. However, no networks of temporal and spatial regulation have yet been defined for these root-specific genes.
Biological functions of plant terpenoids. a Functions of volatile terpenoids (blue arrows) and nonvolatile terpenoids (red arrows) in the interactions of plants with their environment. b Defensive activity of the volatile sesquiterpene, (E)-β-caryophyllene, against infection of Arabidopsis flowers by the microbial pathogen Pseudomonas syringae. Mutants deficient in (E)-β-caryophyllene biosynthesis in the floral stigmatic tissue (tps21) produce lighter seeds post inoculation of flowers with P. syringae. c Antifeedant activity of the semivolatile diterpenoid, rhizathalene, in Arabidopsis roots. Light microscopic pictures of roots of wild-type and rhizathalene biosynthetic mutants (tps08) with and without feeding by Bradysia (fungus gnat) larvae. Increased feeding damage is observed in the absence of the diterpenoid compound (b, c from Huang et al. 2012 [245] and Vaughan et al. 2013 [234], www.plantcell.org, Copyright American Society of Plant Biologists partially modified)
5 Multifunctionality of Plant Terpenoids
Although terpenoids serve important primary functions as photosynthetic pigments (carotenoids), electron carriers (side-chains of ubiquinone and plastoquinone), regulators of growth and development (gibberellins, abscisic acid, strigolactones, brassinosteroids, cytokinins), in protein glycosylation (dolichols), or as elements of membrane structure and function (phytosterols), specialized terpenoid metabolites (covered here), in particular, have been recognized for an array of biological roles. Volatile or semivolatile, low-molecular–weight terpenoids, which include isoprene, monoterpenoids, sesquiterpenoids, and diterpenoids, are implicated in the protection of plants against abiotic stress and in various biotic interactions above- and belowground [236] (Fig. 4a). The substantial emissions of isoprene and monoterpenes from various vascular and nonvascular plants have been associated with the protection against thermal stress. This process is presumably based on an intercalation of the volatile compounds with the photosynthetic membranes and thereby enhances membrane functionality [237–239]. Moreover, transgenic approaches in tobacco and poplar support a role of isoprene in oxidative stress protection [240–243] and are addressed in a separate chapter by Vickers et al.
Volatile terpenoids as constituents of floral scent are implicated in mutualistic interactions with plant pollinators. For instance, choice tests with bumblebees have indicated a role of monoterpenoids emitted by monkeyflowers in pollinator attraction [244]. Nevertheless, distinct evidence for a specific role of terpenoids in pollinator attraction by the use of biosynthetic mutants is still missing, but it can be assumed that attractive effects depend on mixtures of volatiles rather than individual compounds. The notion that floral volatile terpenoids serve multiple functions has been supported by their role in the defense of floral tissues against microbial pathogens. This interaction was demonstrated in flowers of Arabidopsis mutants, which lack the emission of (E)-β-caryophyllene from their stigmatic tissue. The mutant flowers were more susceptible to infection by P. syringae, which resulted in lighter and often misshaped seeds suggesting reduced plant fitness [245] (Fig. 4b). Similar findings were made by Junker et al. [246] demonstrating that floral volatiles play roles in the structuring of bacterial communities that colonize flower petals by providing compound-dependent niches.
Volatile terpenoids also serve important functions as constitutive or pathogen- and herbivore-induced compounds in the defense of photosynthetic tissues. For example, repellent activities have been reported for monoterpene volatiles that are emitted by leaves of Chrysanthemum morifolium and, notably, herbivore-deterrent effects have been observed for isoprene [247, 248]. Furthermore, volatile terpenoids that accumulate in glandular trichomes function as insect repellents as was, for example, found for the activity of sesquiterpenes in trichomes of wild tomato against white flies [249] (Fig. 4a). In conifers, the production of terpenoid oleoresin and terpenoid volatile emissions constitute an important chemical defense system [250]. In a search for resistance factors, the monoterpene (+)-3-carene was found to be associated with resistance of Sitka spruce (Picea sitchensis) to white pine weevil (Pissodes strobi) [251]. Variation of the (+)-3-carene production in resistant and susceptible trees was demonstrated to depend on the copy number of a (+)-3-carene TPS gene, differences in gene transcript and protein levels, and variation in catalytic efficiencies. Similarly, in Arabidopsis, ecotype-specific variation of the herbivore-induced volatiles, (E)-beta-ocimene and (E,E)-alpha-farnesene, is controlled by allelic variation and differences in subcellular targeting of the two terpene synthases, TPS02 and TPS03 [42].
The role of herbivore-induced volatile blends in the attraction of natural enemies of herbivores (Fig. 4a) and at higher trophic levels has been investigated in numerous studies (reviewed by [252, 253]). Work with transgenic Arabidopsis provided strong evidence for the role of volatile terpenes in these interactions [254–256]. However, as indicated for floral scent, the effect of these compounds has to be considered in the context of the entire herbivore-induced volatile blend, and actual fitness benefits to the plant host under natural conditions are still debated [252, 257, 258]. Indirect defense responses mediated by volatile compounds also occur upon insect oviposition [259]. For example, egg deposition on the foliage of European field elm (Ulmus minor) by the elm leaf beetle (Xanthogaleruca luteola) leads to the emission of volatiles including the irregular homoterpene, (E)-4,8-dimethyl-1,3,7-nonatriene (DMNT), which play a role in the attraction of the specialist egg parasitoid, Oomyzus gallerucae [260].
In addition to their function in the interaction with herbivores and their enemies, constitutive and induced volatile mixtures (including volatile terpenes such as homoterpenes) can serve as interspecific, intraspecific, and intraplant “alarm” signals to prime or induce defense responses in neighboring plants or in unattacked tissues of the same plant [261–264]. In these interactions, volatiles may not necessarily need to enter the leaf tissue of the neighboring plant but remain on the leaf surface. This effect was observed for sesquiterpenoids that are emitted by rhododendron leaves and adsorbed on the leaves of birch trees, where they exhibit direct herbivore-repellent activities [265]. Moreover, terpenoids were suggested to be involved in parasitic plant interactions, specifically, in the attraction of the parasitic plant Cuscuta pentagona (dodder) to establish contact with tomato as its host [266]. The molecular mechanism of host plant detection in this response as in other volatile-mediated plant–plant interactions is still poorly understood.
The described functions of volatile terpenoids in aboveground plant defense are complemented by nonvolatile terpenoids. As an example, glycosides of geranyllinalool serve as potent antifeedants in the wild tobacco, Nicotiana obtusifolia [267], and recently detected ent-kaurane–related diterpenoids in maize named kauralexins as well as acidic sesquiterpenes called zealexins function as pathogen-inducible phytoalexins [268, 269]. Similarly, metabolomics studies of Barbarea vulgaris revealed that triterpene saponins contribute to resistance against flea beetle attack [270].
An increased interest in the role of specialized metabolites belowground has shown that terpenoids serve functions similar to those aboveground. Recent studies in Arabidopsis roots discovered semivolatile diterpene hydrocarbons with an unusual tricyclic spiro-hydrindane structure called rhizathalenes [234] (Fig. 4c). These compounds are produced in the root stele, from where they diffuse through the surrounding cell layers to function as local antifeedants by reducing root herbivore damage on these cell layers [234] (Fig. 4c). The role of volatile terpenes in belowground indirect defense has been well established based on studies in maize showing that the sesquiterpene, (E)-β-caryophyllene, which is emitted from roots upon attack by the Western corn root worm Diabrotica virgifera, attracts entomopathogenic nematodes [271, 272]. These findings prompted attempts to engineer (E)-β-caryophyllene production in nonemitting American maize cultivars, which resulted in an increased attraction of nematodes and higher resistance to corn root worm attack [273]. However, constitutive emissions of (E)-β-caryophyllene were found to have additional costs inasmuch as they compromise seed germination, plant growth, and yield [274]. Therefore, more fine-tuned engineering strategies considering herbivore-induced emissions may have to be developed to circumvent these cost effects.
Nonvolatile terpenoids can be exuded from roots into the rhizosphere and the surrounding soil environment where they are involved in different defense responses. Studies using rice mutants convincingly demonstrated that labdane-related diterpenoids named momilactones exhibit allelopathic effects on barnyard grass competitors [275]. Moreover, avenacins, which are triterpene saponins exuded by the roots of oat, are known for their potent activity as phytoalexins [276]. Excitingly, a recent study by Osbourn and colleagues revealed that common triterpene precursors have additional signaling functions in root development. Specifically, it was demonstrated that β-amyrin is involved with determining the patterns of epidermal root hair cells [277]. These findings indicate that the roles of specialized metabolites in biotic interactions and potential “primary” functions become increasingly blurred. Signaling functions have also been demonstrated for the abietane diterpenoid, dehydroabietinal, which is produced at picomolar concentrations in Arabidopsis leaf tissue and serves as a vascular signaling compound and potent activator of systemic acquired resistance [278]. This activity seems to depend on the association of dehydroabietinal with vascular sap proteins.
Finally, it should be noted that strigolactones have become an exciting model for the multifunctionality of small molecules. As carotenoid-derived compounds (reviewed by [279]), strigolactones have important roles as exogenous signals by recruiting arbuscular mycorrhizal fungi in the rhizosphere [280]. Parasitic plants such as Striga lutea (witchweed) eavesdrop on these compounds by using them as germination signals [281]. As internal signals, strigolactones function as growth and developmental hormones that suppress shoot branching [282, 283]. Other processes that involve strigolactone signaling functions include root growth and development, stem elongation, secondary growth, leaf expansion and senescence, and responses to drought and salinity [279, 284]. Rapid progress has been made in understanding the perception of strigolactones but many open questions remain about downstream targets and the role of strigolactone-related compounds [279].
6 Outlook
In the past years, research in terpenoid metabolism has received a boost from developments in synthetic biology to generate engineering platforms for the production of high-value terpenoid products. Production systems in microbes have been developed to result in substantial yields [285], however, engineering of terpenoids in plants still faces challenges because of the complexity of metabolic and regulatory networks. Nevertheless, strategies to avoid metabolic flux competition by targeting biosynthetic modules to different cellular compartments have proved to be promising. Likewise, establishing pathways in specialized cells such as trichomes helps avoid metabolic competition and phytotoxic effects that could negatively affect growth and yield. The discovery of trichome-specific CPTs and TPS enzymes with substrate specificity for cis-prenyl diphosphates most likely will facilitate the engineering efforts in these tissues and provide new gene tools for building synthetic modules. Despite the successful use of distinct organelles such as plastids as “mini” subcellular factories, more efforts need to be made to understand the compartmentalization of the core terpenoid pathway. Especially, additional work should be performed to clarify the putative localization of the MVA pathway and prenyltransferase enzymes in peroxisomes, which would add yet another dimension to the compartmental complexity of terpenoid metabolism in plants. Genomics-based efforts to better understand the regulation of the early terpenoid pathways and terpenoid biosynthetic gene clusters are on their way and will be essential to gain a better understanding of the regulatory networks and epigenetic factors coordinating terpenoid metabolic routes in space and time. Finally, our knowledge of the biological roles of terpenoids is still far from complete. The recent findings of overlapping activities of terpenoids such as strigolactones or triterpenoids in biotic interactions and as internal signals indicate a need to use advanced mutant-based approaches for elucidating the multifunctionality of plant terpenoid compounds.
References
Kubeczka KH (2010) History and sources of essential oil research. In: Baser KHC, Buchbauer G (eds) Handbook of essential oils: science, technology, and applications. CRC Press/Taylor & Francis, Boca Raton, pp 3–38
Zhang FZ, Rodriguez S, Keasling JD (2011) Metabolic engineering of microbial pathways for advanced biofuels production. Curr Opin Biotechnol 22:775–783
Lange BM, Ahkami A (2013) Metabolic engineering of plant monoterpenes, sesquiterpenes and diterpenes-current status and future opportunities. Plant Biotechnol J 11:169–196
Hemmerlin A, Harwood JL, Bach TJ (2012) A raison d’etre for two distinct pathways in the early steps of plant isoprenoid biosynthesis? Prog Lip Res 51:95–148
Hemmerlin A (2013) Post-translational events and modifications regulating plant enzymes involved in isoprenoid precursor biosynthesis. Plant Sci 203:41–54
Vranova E, Coman D, Gruissem W (2013) Network analysis of the MVA and MEP pathways for isoprenoid synthesis. In: Merchant SS (ed) Ann Rev Plant Biol, vol 64, pp 665–700
Vranova E, Coman D, Gruissem W (2012) Structure and dynamics of the isoprenoid pathway network. Mol Plant 5:318–333
Caelles C, Ferrer A, Balcells L, Hegardt FG, Boronat A (1989) Isolation and structural characterization of a cDNA-encoding Arabidopsis thaliana 3-hydroxy-3-methylglutaryl coenzyme A reductase. Plant Mol Biol 13:627–638
Enjuto M, Balcells L, Campos N, Caelles C, Arro M, Boronat A (1994) Arabidopsis thaliana contains 2 differentially expressed 3-hydroxy-3-methylglutaryl-CoA reductase genes, which encode microsomal forms of the enzyme. Proc Natl Acad Sci USA 91:927–931
Campos N, Boronat A (1995) Targeting and topology in the membrane of plant 3-hydroxy-3-methylglutaryl coenzyme a reductase. Plant Cell 7:2163–2174
Denbow CJ, Lang S, Cramer CL (1995) Targeting and membrane orientation of tomato 3-hydroxy-3-methylglutaryl coenzyme A reductases. Plant Physiol 108:144
Re EB, Brugger S, Learned M (1997) Genetic and biochemical analysis of the transmembrane domain of Arabidopsis 3-hydroxy-3-methylglutaryl coenzyme A reductase. J Cell Biochem 65:443–459
Vollack KU, Dittrich B, Ferrer A, Boronat A, Bach TJ (1994) Two radish genes for 3-hydroxy-3-methylglutaryl-CoA reductase isozymes complement mevalonate auxotrophy in a yeast mutant and yield membrane-bound active enzyme. J Plant Physiol 143:479–487
Re EB, Jones D, Learned RM (1995) Coexpression of native and introduced genes reveals cryptic regulation of HMG CoA reductase expression in Arabidopsis. Plant J 7:771–784
Holmberg N, Harker M, Wallace AD, Clayton JC, Gibbard CL, Safford R (2003) Co-expression of N-terminal truncated 3-hydroxy-3-methylglutaryl CoA reductase and C24-sterol methyltransferase type 1 in transgenic tobacco enhances carbon flux towards end-product sterols. Plant J 36:12–20
Ohyama K, Suzuki M, Masuda K, Yoshida S, Muranaka T (2007) Chemical phenotypes of the hmg1 and hmg2 mutants of Arabidopsis demonstrate the in-planta role of HMG-CoA reductase in triterpene biosynthesis. Chem Pharm Bull (Tokyo) 55:1518–1521
Suzuki M, Kamide Y, Nagata N, Seki H, Ohyama K, Kato H, Masuda K, Sato S, Kato T, Tabata S, Yoshida S, Muranaka T (2004) Loss of function of 3-hydroxy-3-methylglutaryl coenzyme A reductase 1 (HMG1) in Arabidopsis leads to dwarfing, early senescence and male sterility, and reduced sterol levels. Plant J 37:750–761
Manzano D, Fernandez-Busquets X, Schaller H, Gonzalez V, Boronat A, Arro M, Ferrer A (2004) The metabolic imbalance underlying lesion formation in Arabidopsis thaliana overexpressing farnesyl diphosphate synthase (isoform 1S) leads to oxidative stress and is triggered by the developmental decline of endogenous HMGR activity. Planta 219:982–992
Chappell J, Vonlanken C, Vogeli U (1991) Elicitor-inducible 3-hydroxy-3-methylglutaryl coenzyme A reductase activity is required for sesquiterpene accumulation in tobacco cell-suspension cultures. Plant Physiol 97:693–698
Chappell J, Wolf F, Proulx J, Cuellar R, Saunders C (1995) Is the reaction catalyzed by 3-hydroxy-3-methylglutaryl coenzyme A reductase a rate-limiting step for isoprenoid biosynthesis in plants? Plant Physiol 109:1337–1343
Choi D, Bostock RM, Avdiushko S, Hildebrand DF (1994) Lipid-derived signals that discriminate wound-responsive and pathogen-responsive isoprenoid pathways in plants—methyl jasmonate and the fungal elicitor arachidonic acid induce different 3-hydroxy-3-methylglutaryl coenzyme A reductase genes and antimicrobial isoprenoids in Solanum tuberosum L. Proc Natl Acad Sci USA 91:2329–2333
Lichtenthaler HK (1998) The plant 1-deoxy-d-xylulose-5-phosphate pathway for biosynthesis of isoprenoids. Fett-Lipid 100:128–138
Rohmer M (1999) The discovery of a mevalonate-independent pathway for isoprenoid biosynthesis in bacteria, algae and higher plants. Nat Prod Rep 16:565–574
Boucher Y, Doolittle WF (2000) The role of lateral gene transfer in the evolution of isoprenoid biosynthesis pathways. Mol Microbiol 37:703–716
Lange BM, Rujan T, Martin W, Croteau R (2000) Isoprenoid biosynthesis: the evolution of two ancient and distinct pathways across genomes. Proc Natl Acad Sci USA 97:13172–13177
Bouvier F, d’Harlingue A, Suire C, Backhaus RA, Camara B (1998) Dedicated roles of plastid transketolases during the early onset of isoprenoid biogenesis in pepper fruits. Plant Physiol 117:1423–1431
Lange BM, Wildung MR, McCaskill D, Croteau R (1998) A family of transketolases that directs isoprenoid biosynthesis via a mevalonate-independent pathway. Proc Natl Acad Sci USA 95:2100–2104
Chahed K, Oudin A, Guivarc’h N, Hamdi S, Chenieux JC, Rideau M, Clastre M (2000) l-Deoxy-d-xylulose 5-phosphate synthase from periwinkle: cDNA identification and induced gene expression in terpenoid indole alkaloid-producing cells. Plant Physiol Biochem 38:559–566
Lois LM, Rodriguez-Concepcion M, Gallego F, Campos N, Boronat A (2000) Carotenoid biosynthesis during tomato fruit development: regulatory role of 1-deoxy-d-xylulose 5-phosphate synthase. Plant J 22:503–513
Walter MH, Fester T, Strack D (2000) Arbuscular mycorrhizal fungi induce the non-mevalonate methylerythritol phosphate pathway of isoprenoid biosynthesis correlated with accumulation of the ‘yellow pigment’ and other apocarotenoids. Plant J 21:571–578
Estevez JM, Cantero A, Reindl A, Reichler S, Leon P (2001) 1-Deoxy-d-xylulose-5-phosphate synthase, a limiting enzyme for plastidic isoprenoid biosynthesis in plants. J Biol Chem 276:22901–22909
Mandel MA, Feldmann KA, HerreraEstrella L, RochaSosa M, Leon P (1996) CLA1, a novel gene required for chloroplast development, is highly conserved in evolution. Plant J 9:649–658
Estevez JM, Cantero A, Romero C, Kawaide H, Jimenez LF, Kuzuyama T, Seto H, Kamiya Y, Leon P (2000) Analysis of the expression of CLA1, a gene that encodes the 1-deoxyxylulose 5-phosphate synthase of the 2-C-methyl-d-erythritol-4-phosphate pathway in Arabidopsis. Plant Physiol 124:95–103
Araki N, Kusumi K, Masamoto K, Niwa Y, Iba K (2000) Temperature-sensitive Arabidopsis mutant defective in 1-deoxy-d-xylulose 5-phosphate synthase within the plastid non-mevalonate pathway of isoprenoid biosynthesis. Physiol Plant 108:19–24
Schwender J, Muller C, Zeidler J, Lichlenthaler HK (1999) Cloning and heterologous expression of a cDNA encoding 1-deoxy-d-xylulose-5-phosphate reductoisomerase of Arabidopsis thaliana. FEBS Lett 455:140–144
Carretero-Paulet L, Ahumada I, Cunillera N, Rodriguez-Concepcion M, Ferrer A, Boronat A, Campos N (2002) Expression and molecular analysis of the Arabidopsis DXR gene encoding 1-deoxy-d-xylulose 5-phosphate reductoisomerase, the first committed enzyme of the 2-C-methyl-d-erythritol 4-phosphate pathway. Plant Physiol 129:1581–1591
Kuzuyama T, Shimizu T, Takahashi S, Seto H (1998) Fosmidomycin, a specific inhibitor of 1-deoxy-d-xylulose 5-phosphate reductoisomerase in the nonmevalonate pathway for terpenoid biosynthesis. Tetrahedron Lett 39:7913–7916
Jomaa H, Wiesner J, Sanderbrand S, Altincicek B, Weidemeyer C, Hintz M, Turbachova I, Eberl M, Zeidler J, Lichtenthaler HK, Soldati D, Beck E (1999) Inhibitors of the nonmevalonate pathway of isoprenoid biosynthesis as antimalarial drugs. Science 285:1573–1576
Steinbacher S, Kaiser J, Eisenreich W, Huber R, Bacher A, Rohdich F (2003) Structural basis of fosmidomycin action revealed by the complex with 2-C-methyl-d-erythritol 4-phosphate synthase (IspC)—implications for the catalytic mechanism and anti-malaria drug development. J Biol Chem 278:18401–18407
Zeidler J, Schwender J, Muller C, Wiesner J, Weidemeyer C, Beck E, Jomaa H, Lichtenthaler HK (1998) Inhibition of the non-mevalonate 1-deoxy-d-xylulose-5-phosphate pathway of plant isoprenoid biosynthesis by fosmidomycin. Z fur Naturforsch C- J Biosci 53:980–986
Rodriguez-Concepcion M, Ahumada I, Diez-Juez E, Sauret-Gueto S, Lois LM, Gallego F, Carretero-Paulet L, Campos N, Boronat A (2001) 1-Deoxy-d-xylulose 5-phosphate reductoisomerase and plastid isoprenoid biosynthesis during tomato fruit ripening. Plant J 27:213–222
Huang MS, Abel C, Sohrabi R, Petri J, Haupt I, Cosimano J, Gershenzon J, Tholl D (2010) Variation of herbivore-induced volatile terpenes among Arabidopsis ecotypes depends on allelic differences and subcellular targeting of two terpene synthases, TPS02 and TPS03. Plant Physiol 153:1293–1310
Xing SF, Miao J, Li SA, Qin GJ, Tang S, Li HN, Gu HY, Qu LJ (2010) Disruption of the 1-deoxy-d-xylulose-5-phosphate reductoisomerase (DXR) gene results in albino, dwarf and defects in trichome initiation and stomata closure in Arabidopsis. Cell Res 20:688–700
Rohdich F, Wungsintaweekul J, Fellermeier M, Sagner S, Herz S, Kis K, Eisenreich W, Bacher A, Zenk MH (1999) Cytidine 5’-triphosphate-dependent biosynthesis of isoprenoids: YgbP protein of Escherichia coli catalyzes the formation of 4-diphosphocytidyl-2-C-methylerythritol. Proc Natl Acad Sci USA 96:11758–11763
Rohdich F, Wungsintaweekul J, Eisenreich W, Richter G, Schuhr CA, Hecht S, Zenk MH, Bacher A (2000) Biosynthesis of terpenoids: 4-Diphosphocytidyl-2C-methyl-d-erythritol synthase of Arabidopsis thaliana. Proc Natl Acad Sci USA 97:6451–6456
Lange BM, Croteau R (1999) Isopentenyl diphosphate biosynthesis via a mevalonate-independent pathway: Isopentenyl monophosphate kinase catalyzes the terminal enzymatic step. Proc Natl Acad Sci USA 96:13714–13719
Luttgen H, Rohdich F, Herz S, Wungsintaweekul J, Hecht S, Schuhr CA, Fellermeier M, Sagner S, Zenk MH, Bacher A, Eisenreich W (2000) Biosynthesis of terpenoids: YchB protein of Escherichia coli phosphorylates the 2-hydroxy group of 4-diphosphocytidyl-2C-methyl-d-erythritol. Proc Natl Acad Sci USA 97:1062–1067
Rohdich F, Wungsintaweekul J, Luttgen H, Fischer M, Eisenreich W, Schuhr CA, Fellermeier M, Schramek N, Zenk MH, Bacher A (2000) Biosynthesis of terpenoids: 4-diphosphocytidyl-2-C-methyl-d-erythritol kinase from tomato. Proc Natl Acad Sci USA 97:8251–8256
Rohdich F, Hecht S, Gärtner K, Adam P, Krieger C, Amslinger S, Arigoni D, Bacher A, Eisenreich W (2002) Studies on the nonmevalonate terpene biosynthetic pathway: metabolic role of IspH (LytB) protein. Proc Natl Acad Sci USA 99:1158–1163
Rohdich F, Zepeck F, Adam P, Hecht S, Kaiser J, Laupitz R, Grawert T, Amslinger S, Eisenreich W, Bacher A, Arigoni D (2003) The deoxyxylulose phosphate pathway of isoprenoid biosynthesis: studies on the mechanisms of the reactions catalyzed by IspG and IspH protein. Proc Natl Acad Sci USA 100:1586–1591
Tritsch D, Hemmerlin A, Bach TJ, Rohmer M (2010) Plant isoprenoid biosynthesis via the MEP pathway: in vivo IPP/DMAPP ratio produced by (E)-4-hydroxy-3-methylbut-2-enyl diphosphate reductase in tobacco BY-2 cell cultures. FEBS Lett 584:129–134
Hsieh MH, Goodman HM (2006) Functional evidence for the involvement of Arabidopsis IspF homolog in the nonmevalonate pathway of plastid isoprenoid biosynthesis. Planta 223:779–784
Hsieh MH, Chang CY, Hsu SJ, Chen JJ (2008) Chloroplast localization of methylerythritol 4-phosphate pathway enzymes and regulation of mitochondrial genes in ispD and ispE albino mutants in Arabidopsis. Plant Mol Biol 66:663–673
Gutierrez-Nava MDL, Gillmor CS, Jimenez LF, Guevara-Garcia A, Leon P (2004) Chloroplast biogenesis genes act cell and noncell autonomously in early chloroplast development. Plant Physiol 135:471–482
Hsieh MH, Goodman HM (2005) The Arabidopsis IspH homolog is involved in the plastid nonmevalonate pathway of isoprenoid biosynthesis. Plant Physiol 138:641–653
Gil MJ, Coego A, Mauch-Mani B, Jorda L, Vera P (2005) The Arabidopsis csb3 mutant reveals a regulatory link between salicylic acid-mediated disease resistance and the methyl-erythritol 4-phosphate pathway. Plant J 44:155–166
Wang H, Nagegowda DA, Rawat R, Bouvier-Nave P, Guo DJ, Bach TJ, Chye ML (2012) Overexpression of Brassica juncea wild-type and mutant HMG-CoA synthase 1 in Arabidopsis up-regulates genes in sterol biosynthesis and enhances sterol production and stress tolerance. Plant Biotechnol J 10:31–42
Alex D, Bach TJ, Chye ML (2000) Expression of Brassica juncea 3-hydroxy-3-methylglutaryl CoA synthase is developmentally regulated and stress-responsive. Plant J 22:415–426
Nagegowda DA, Ramalingam S, Hemmerlin A, Bach TJ, Chye ML (2005) Brassica juncea HMG-CoA synthase: localization of mRNA and protein. Planta 221:844–856
Ishiguro S, Nishimori Y, Yamada M, Saito H, Suzuki T, Nakagawa T, Miyake H, Okada K, Nakamura K (2010) The Arabidopsis FLAKY POLLEN1 gene encodes a 3-hydroxy-3-methylglutaryl-coenzyme A synthase required for development of tapetum-specific organelles and fertility of pollen grains. Plant Cell Physiol 51:896–911
Choi D, Ward BL, Bostock RM (1992) Differential induction and suppression of potato 3-hydroxy-3-methylglutaryl coenezyme A reductase genes in response to Phytophthora infestans and to its elicitor arachidonic acid. Plant Cell 4:1333–1344
Rodriguez-Concepcion M, Gruissem W (1999) Arachidonic acid alters tomato HMG expression and fruit growth and induces 3-hydroxy-3-methylglutaryl coenzyme A reductase-independent lycopene accumulation. Plant Physiol 119:41–48
Suzuki H, Xia YJ, Cameron R, Shadle G, Blount J, Lamb C, Dixon RA (2004) Signals for local and systemic responses of plants to pathogen attack. J Exp Bot 55:169–179
Suzuki M, Nakagawa S, Kamide Y, Kobayashi K, Ohyama K, Hashinokuchi H, Kiuchi R, Saito K, Muranaka T, Nagata N (2009) Complete blockage of the mevalonate pathway results in male gametophyte lethality. J Exp Biol 60:2055–2064
Walter MH, Hans J, Strack D (2002) Two distantly related genes encoding 1-deoxy-d-xylulose 5-phosphate synthases: differential regulation in shoots and apocarotenoid-accumulating mycorrhizal roots. Plant J 31:243–254
Paetzold H, Garms S, Bartram S, Wieczorek J, Uros-Gracia EM, Rodriguez-Concepcion M, Boland W, Strack D, Hause B, Walter MH (2010) The isogene 1-deoxy-d-xylulose 5-phosphate synthase 2 controls isoprenoid profiles, precursor pathway allocation, and density of tomato trichomes. Mol Plant 3:904–916
Brooker JD, Russell DW (1975) Properties of microsomal 3-hydroxy-3-methylglutaryl coenzyme A reductase from Pisum sativum seedlings. Arch Biochem Biophys 167:723–729
Soto G, Stritzler M, Lisi C, Alleva K, Pagano ME, Ardila F, Mozzicafreddo M, Cuccioloni M, Angeletti M, Ayub ND (2011) Acetoacetyl-CoA thiolase regulates the mevalonate pathway during abiotic stress adaptation. J Exp Bot 62:5699–5711
Schulte AE, van der Heijden R, Verpoorte R (2000) Purification and characterization of mevalonate kinase from suspension-cultured cells of Catharanthus roseus (L.) G. Don. Arch Biochem Biophys 378:287–298
Banerjee A, Wu Y, Banerjee R, Li Y, Yan HG, Sharkey TD (2013) Feedback inhibition of deoxy-d-xylulose-5-phosphate synthase regulates the methylerythritol 4-phosphate pathway. J Biol Chem 288:16926–16936
Ghirardo A, Wright LP, Bi Z, Rosenkranz M, Pulido P, Rodriguez-Concepcion M, Niinemets U, Brueggemann N, Gershenzon J, Schnitzler J-P (2014) Metabolic flux analysis of plastidic isoprenoid biosynthesis in poplar leaves emitting and nonemitting isoprene. Plant Physiol 165:37–51
Liao P, Wang H, Wang M, Hsiao A-S, Bach TJ, Chye M-L (2014) Transgenic tobacco overexpressing Brassica juncea HMG-CoA Synthase 1 shows increased plant growth, pod size, and seed yield. Plos One 9:e98264
Jin HN, Song ZH, Nikolau BJ (2012) Reverse genetic characterization of two paralogous acetoacetyl CoA thiolase genes in Arabidopsis reveals their importance in plant growth and development. Plant J 70:1015–1032
Holmberg N, Harker M, Gibbard CL, Wallace AD, Clayton JC, Rawlins S, Hellyer A, Safford R (2002) Sterol C-24 methyltransferase type 1 controls the flux of carbon into sterol biosynthesis in tobacco seed. Plant Physiol 130:303–311
Wentzinger LF, Bach TJ, Hartmann MA (2002) Inhibition of squalene synthase and squalene epoxidase in tobacco cells triggers an up-regulation of 3-hydroxy-3-methylglutaryl coenzyme A reductase. Plant Physiol 130:334–346
Ahn CS, Pai HS (2008) Physiological function of ispE, a plastid MEP pathway gene for isoprenoid biosynthesis, in organelle biogenesis and cell morphogenesis in Nicotiana benthamiana. Plant Mol Biol 66:503–517
Jung KH, Lee J, Dardick C, Seo YS, Cao P, Canlas P, Phetsom J, Xu X, Ouyang S, An K, Cho YJ, Lee GC, Lee Y, An G, Ronald PC (2008) Identification and functional analysis of light-responsive unique genes and gene family members in rice. PLoS Genet 4
Wille A, Zimmermann P, Vranova E, Furholz A, Laule O, Bleuler S, Hennig L, Prelic A, von Rohr P, Thiele L, Zitzler E, Gruissem W, Buhlmann P (2004) Sparse graphical Gaussian modeling of the isoprenoid gene network in Arabidopsis thaliana. Genome Biol 5
Ghassemian M, Lutes J, Tepperman JM, Chang HS, Zhu T, Wang X, Quail PH, Lange BM (2006) Integrative analysis of transcript and metabolite profiling data sets to evaluate the regulation of biochemical pathways during photomorphogenesis. Arch Biochem Biophys 448:45–59
Meier S, Tzfadia O, Vallabhaneni R, Gehring C, Wurtzel ET (2011) A transcriptional analysis of carotenoid, chlorophyll and plastidial isoprenoid biosynthesis genes during development and osmotic stress responses in Arabidopsis thaliana. BMC Syst Biol 5
Flores-Perez U, Perez-Gila J, Closa M, Wright LP, Botella-Pavia P, Phillips MA, Ferrer A, Gershenzon J, Rodriguez-Concepcion M (2010) PLEIOTROPIC REGULATORY LOCUS 1 (PRL1) integrates the regulation of sugar responses with isoprenoid metabolism in Arabidopsis. Mol Plant 3:101–112
Dale S, Arro M, Becerra B, Morrice NG, Boronat A, Hardie DG, Ferrer A (1995) Bacterial expression of the catalytic domain of 3-hydroxy-3-methylglutaryl-CoA reductase (isoform HMGR1) from Arabidopsis thaliana, and its inactivation by phosphorylation at Ser577 by Brassica oleracea 3-hydroxy-3-methylglutaryl-CoA reductase kinase. Eur J Biochem 233:506–513
Leivar P, Antolin-Llovera M, Ferrero S, Closa M, Arro M, Ferrer A, Boronat A, Camposa N (2011) Multilevel control of Arabidopsis 3-hydroxy-3-methylglutaryl coenzyme A reductase by protein phosphatase 2A. Plant Cell 23:1494–1511
Yoshioka H, Miyabe M, Hayakawa Y, Doke N (1996) Expression of genes for phenylalanine ammonia-lyase and 3-hydroxy-3-methylglutaryl CoA reductase in aged potato tubers infected with Phytophthora infestans. Plant Cell Physiol 37:81–90
Nieto B, Fores O, Arro M, Ferrer A (2009) Arabidopsis 3-hydroxy-3-methylglutaryl-CoA reductase is regulated at the post-translational level in response to alterations of the sphingolipid and the sterol biosynthetic pathways. Phytochemistry 70:53–59
Kang JH, McRoberts J, Shi F, Moreno JE, Jones AD, Howe GA (2014) The flavonoid biosynthetic enzyme chalcone isomerase modulates terpenoid production in glandular trichomes of tomato. Plant Physiol 164:1161–1174
Pourcel L, Irani NG, Koo AJK, Bohorquez-Restrepo A, Howe GA, Grotewold E (2013) A chemical complementation approach reveals genes and interactions of flavonoids with other pathways. Plant J 74:383–397
Sahu NK, Balbhadra SS, Choudhary J, Kohli DV (2012) Exploring pharmacological significance of chalcone scaffold: a review. Curr Med Chem 19:209–225
Saslowsky DE, Warek U, Winkel BSJ (2005) Nuclear localization of flavonoid enzymes in Arabidopsis. J Biol Chem 280:23735–23740
Ben Zvi MM, Shklarman E, Masci T, Kalev H, Debener T, Shafir S, Ovadis M, Vainstein A (2012) PAP1 transcription factor enhances production of phenylpropanoid and terpenoid scent compounds in rose flowers. New Phytol 195:335–345
Cordoba E, Salmi M, Leon P (2009) Unravelling the regulatory mechanisms that modulate the MEP pathway in higher plants. J Exp Bot 60:2933–2943
Rodriguez-Concepcion M, Fores O, Martinez-Garcia JF, Gonzalez V, Phillips MA, Ferrer A, Boronat A (2004) Distinct light-mediated pathways regulate the biosynthesis and exchange of isoprenoid precursors during Arabidopsis seedling development. Plant Cell 16:144–156
Toledo-Ortiz G, Huq E, Rodriguez-Concepcion M (2010) Direct regulation of phytoene synthase gene expression and carotenoid biosynthesis by phytochrome-interacting factors. Proc Natl Acad Sci USA 107:11626–11631
Wiberley AE, Donohue AR, Westphal MM, Sharkey TD (2009) Regulation of isoprene emission from poplar leaves throughout a day. Plant Cell Environ 32:939–947
Mongelard G, Seemann M, Boisson AM, Rohmer M, Bligny R, Rivasseau C (2011) Measurement of carbon flux through the MEP pathway for isoprenoid synthesis by P-31-NMR spectroscopy after specific inhibition of 2-C-methyl-d-erythritol 2,4-cyclodiphosphate reductase. Effect of light and temperature. Plant Cell Environ 34:1241–1247
Kim YJ, Lee OR, Oh JY, Jang MG, Yang DC (2014) Functional analysis of 3-hydroxy-3-methylglutaryl coenzyme A reductase encoding genes in triterpene saponin-producing ginseng. Plant Physiol 165:373–387
Mannen K, Matsumoto T, Takahashi S, Yamaguchi Y, Tsukagoshi M, Sano R, Suzuki H, Sakurai N, Shibata D, Koyama T, Nakayama T (2014) Coordinated transcriptional regulation of isopentenyl diphosphate biosynthetic pathway enzymes in plastids by phytochrome-interacting factor 5. Biochem Biophys Res Commun 443:768–774
Flores-Perez U, Sauret-Gueto S, Gas E, Jarvis P, Rodriguez-Concepcion M (2008) A mutant impaired in the production of plastome-encoded proteins uncovers a mechanism for the homeostasis of isoprenoid biosynthetic enzymes in Arabidopsis plastids. Plant Cell 20:1303–1315
Fukushima A, Kusano M, Nakamichi N, Kobayashi M, Hayashi N, Sakakibara H, Mizuno T, Saito K (2009) Impact of clock-associated Arabidopsis pseudo-response regulators in metabolic coordination. Proc Natl Acad Sci USA 106:7251–7256
Dudareva N, Andersson S, Orlova I, Gatto N, Reichelt M, Rhodes D, Boland W, Gershenzon J (2005) The nonmevalonate pathway supports both monoterpene and sesquiterpene formation in snapdragon flowers. Proc Natl Acad Sci USA 102:933–938
Lemaire SD, Guillon B, Le Marechal P, Keryer E, Miginiac-Maslow M, Decottignies P (2004) New thioredoxin targets in the unicellular photosynthetic eukaryote Chlamydomonas reinhardtii. Proc Natl Acad Sci USA 101:7475–7480
Balmer Y, Koller A, del Val G, Manieri W, Schurmann P, Buchanan BB (2003) Proteomics gives insight into the regulatory function of chloroplast thioredoxins. Proc Natl Acad Sci USA 100:370–375
Seemann M, Bui BTS, Wolff M, Miginlac-Maslow M, Rohmer M (2006) Isoprenoid biosynthesis in plant chloroplasts via the MEP pathway: direct thylakoid/ferredoxin-dependent photoreduction of GcpE/IspG. FEBS Lett 580:1547–1552
Vogeli U, Chappell J (1988) Induction of sesquiterpene cyclase and suppression of squalene synthase activities in plant cell cultures treated with fungal elicitor. Plant Physiol 88:1291–1296
Kim CY, Zhang SQ (2004) Activation of a mitogen-activated protein kinase cascade induces WRKY family of transcription factors and defense genes in tobacco. Plant J 38:142–151
Jin HL, Liu YD, Yang KY, Kim CY, Baker B, Zhang SQ (2003) Function of a mitogen-activated protein kinase pathway in N gene-mediated resistance in tobacco. Plant J 33:719–731
Kevei Z, Lougnon G, Mergaert P, Horvath GV, Kereszt A, Jayaraman D, Zaman N, Marcel F, Regulski K, Kiss GB, Kondorosi A, Endre G, Kondorosi E, Ane JM (2007) 3-hydroxy-3-methylglutaryl coenzyme A reductase1 interacts with NORK and is crucial for nodulation in Medicago truncatula. Plant Cell 19:3974–3989
Sapir-Mir M, Mett A, Belausov E, Tal-Meshulam S, Frydman A, Gidoni D, Eyal Y (2008) Peroxisomal localization of Arabidopsis isopentenyl diphosphate isomerases suggests that part of the plant isoprenoid mevalonic acid pathway is compartmentalized to peroxisomes. Plant Physiol 148:1219–1228
Simkin AJ, Guirimand G, Papon N, Courdavault V, Thabet I, Ginis O, Bouzid S, Giglioli-Guivarc’h N, Clastre M (2011) Peroxisomal localisation of the final steps of the mevalonic acid pathway in planta. Planta 234:903–914
Guirimand G, Guihur A, Phillips MA, Oudin A, Glevarec G, Melin C, Papon N, Clastre M, St-Pierre B, Rodriguez-Concepcion M, Burlat V, Courdavault V (2012) A single gene encodes isopentenyl diphosphate isomerase isoforms targeted to plastids, mitochondria and peroxisomes in Catharanthus roseus. Plant Mol Biol 79:443–459
Lichtenthaler HK (1999) The 1-deoxy-d-xylulose-5-phosphate pathway of isoprenoid biosynthesis in plants. Annu Rev Plant Physiol Plant Mol Biol 50:47–65
Hemmerlin A, Hoeffler JF, Meyer O, Tritsch D, Kagan IA, Grosdemange-Billiard C, Rohmer M, Bach TJ (2003) Cross-talk between the cytosolic mevalonate and the plastidial methylerythritol phosphate pathways in tobacco bright yellow-2 cells. J Biol Chem 278:26666–26676
Laule O, Furholz A, Chang HS, Zhu T, Wang X, Heifetz PB, Gruissem W, Lange BM (2003) Crosstalk between cytosolic and plastidial pathways of isoprenoid biosynthesis in Arabidopsis thaliana. Proc Natl Acad Sci USA 100:6866–6871
Bartram S, Jux A, Gleixner G, Boland W (2006) Dynamic pathway allocation in early terpenoid biosynthesis of stress-induced lima bean leaves. Phytochemistry 67:1661–1672
Chaurasiya ND, Sangwan NS, Sabir F, Misra L, Sangwan RS (2012) Withanolide biosynthesis recruits both mevalonate and DOXP pathways of isoprenogenesis in Ashwagandha Withania somnifera L. (Dunal). Plant Cell Rep 31:1889–1897
Woelwer-Rieck U, May B, Lankes C, Wuest M (2014) Methylerythritol and mevalonate pathway contributions to biosynthesis of mono-, sesqui-, and diterpenes in glandular trichomes and leaves of Stevia rebaudiana Bertoni. J Agric Food Chem 62:2428–2435
Zhao S, Wang L, Liu L, Liang Y, Sun Y, Wu J (2014) Both the mevalonate and the non-mevalonate pathways are involved in ginsenoside biosynthesis. Plant Cell Rep 33:393–400
Opitz S, Nes WD, Gershenzon J (2014) Both methylerythritol phosphate and mevalonate pathways contribute to biosynthesis of each of the major isoprenoid classes in young cotton seedlings. Phytochemistry 98:110–119
Bick JA, Lange BM (2003) Metabolic cross talk between cytosolic and plastidial pathways of isoprenoid biosynthesis: unidirectional transport of intermediates across the chloroplast envelope membrane. Arch Biochem Biophys 415:146–154
Flügge UI, Gao W (2005) Transport of isoprenoid intermediates across chloroplast envelope membranes. Plant Biol 7:91–97
Gutensohn M, Orlova I, Nguyen TTH, Davidovich-Rikanati R, Ferruzzi MG, Sitrit Y, Lewinsohn E, Pichersky E, Dudareva N (2013) Cytosolic monoterpene biosynthesis is supported by plastid-generated geranyl diphosphate substrate in transgenic tomato fruits. Plant J 75:351–363
May B, Lange BM, Wuest M (2013) Biosynthesis of sesquiterpenes in grape berry exocarp of Vitis vinifera L.: evidence for a transport of farnesyl diphosphate precursors from plastids to the cytosol. Phytochemistry 95:135–144
Wu SQ, Schalk M, Clark A, Miles RB, Coates R, Chappell J (2006) Redirection of cytosolic or plastidic isoprenoid precursors elevates terpene production in plants. Nat Biotechnol 24:1441–1447
Wu S, Jiang Z, Kempinski C, Nybo SE, Husodo S, Williams R, Chappell J (2012) Engineering triterpene metabolism in tobacco. Planta 236:867–877
Kumar S, Hahn FM, Baidoo E, Kahlon TS, Wood DF, McMahan CM, Cornish K, Keasling JD, Daniell H, Whalen MC (2012) Remodeling the isoprenoid pathway in tobacco by expressing the cytoplasmic mevalonate pathway in chloroplasts. Metab Eng 14:19–28
Huchelmann A, Gastaldo C, Veinante M, Zeng Y, Heintz D, Tritsch D, Schaller H, Rohmer M, Bach TJ, Hemmerlin A (2014) S-Carvone suppresses cellulase-induced capsidiol production in Nicotiana tabacum by interfering with protein isoprenylation. Plant Physiol 164:935–950
Verbitskiy D, Zehrmann A, van der Merwe JA, Brennicke A, Takenaka M (2010) The PPR protein encoded by the Lovastatin Insensitive 1 gene is involved in RNA editing at three sites in mitochondria of Arabidopsis thaliana. Plant J 61:446–455
Tang JW, Kobayashi K, Suzuki M, Matsumoto S, Muranaka T (2010) The mitochondrial PPR protein LOVASTATIN INSENSITIVE 1 plays regulatory roles in cytosolic and plastidial isoprenoid biosynthesis through RNA editing. Plant J 61:456–466
Xiao YM, Savchenko T, Baidoo EEK, Chehab WE, Hayden DM, Tolstikov V, Corwin JA, Kliebenstein DJ, Keasling JD, Dehesh K (2012) Retrograde signaling by the plastidial metabolite MEcPP regulates expression of nuclear stress-response genes. Cell 149:1525–1535
Phillips MA, D’Auria JC, Gershenzon J, Pichersky E (2008) The Arabidopsis thaliana type I isopentenyl diphosphate isomerases are targeted to multiple subcellular compartments and have overlapping functions in isoprenoid biosynthesis. Plant Cell 20:677–696
Kharel Y, Koyama T (2003) Molecular analysis of cis-prenyl chain elongating enzymes. Nat Prod Rep 20:111–118
Burke C, Croteau R (2002) Interaction with the small subunit of geranyl diphosphate synthase modifies the chain length specificity of geranylgeranyl diphosphate synthase to produce geranyl diphosphate. J Biol Chem 277:3141–3149
Schmidt A, Gershenzon J (2008) Cloning and characterization of two different types of geranyl diphosphate synthases from Norway spruce (Picea abies). Phytochemistry 69:49–57
Chang TH, Hsieh FL, Ko TP, Teng KH, Liang PH, Wang AHJ (2010) Structure of a heterotetrameric geranyl pyrophosphate synthase from mint (Mentha piperita) reveals intersubunit regulation. Plant Cell 22:454–467
Rai A, Smita SS, Singh AK, Shanker K, Nagegowda DA (2013) Heteromeric and homomeric geranyl diphosphate synthases from Catharanthus roseus and their role in monoterpene indole alkaloid biosynthesis. Mol Plant 6:1531–1549
Hsiao YY, Jeng MF, Tsai WC, Chuang YC, Li CY, Wu TS, Kuoh CS, Chen WH, Chen HH (2008) A novel homodimeric geranyl diphosphate synthase from the orchid Phalaenopsis bellina lacking a DD(X)(2-4)D motif. Plant J 55:719–733
Burke CC, Wildung MR, Croteau R (1999) Geranyl diphosphate synthase: cloning, expression, and characterization of this prenyltransferase as a heterodimer. Proc Natl Acad Sci USA 96:13062–13067
Tholl D, Kish CM, Orlova I, Sherman D, Gershenzon J, Pichersky E, Dudareva N (2004) Formation of monoterpenes in Antirrhinum majus and Clarkia breweri flowers involves heterodimeric geranyl diphosphate synthases. Plant Cell 16:977–992
Wang GD, Dixon RA (2009) Heterodimeric geranyl(geranyl)diphosphate synthase from hop (Humulus lupulus) and the evolution of monoterpene biosynthesis. Proc Natl Acad Sci USA 106:9914–9919
Orlova I, Nagegowda DA, Kish CM, Gutensohn M, Maeda H, Varbanova M, Fridman E, Yamaguchi S, Hanada A, Kamiya Y, Krichevsky A, Citovsky V, Pichersky E, Dudareva N (2009) The small subunit of snapdragon geranyl diphosphate synthase modifies the chain length specificity of tobacco geranylgeranyl diphosphate synthase in planta. Plant Cell 21:4002–4017
van Schie CCN, Ament K, Schmidt A, Lange T, Haring MA, Schuurink RC (2007) Geranyl diphosphate synthase is required for biosynthesis of gibberellins. Plant J 52:752–762
Schmidt A, Wachtler B, Temp U, Krekling T, Seguin A, Gershenzon J (2010) A bifunctional geranyl and geranylgeranyl diphosphate synthase is involved in terpene oleoresin formation in Picea abies. Plant Physiol 152:639–655
Bouvier F, Suire C, d’Harlingue A, Backhaus RA, Camara B (2000) Molecular cloning of geranyl diphosphate synthase and compartmentation of monoterpene synthesis in plant cells. Plant J 24:241–252
Hsieh F-L, Chang T-H, Ko T-P, Wang AHJ (2011) Structure and mechanism of an Arabidopsis medium/long-chain-length prenyl pyrophosphate synthase. Plant Physiol 155:1079–1090
Cunillera N, Arro M, Delourme D, Karst F, Boronat A, Ferrer A (1996) Arabidopsis thaliana contains two differentially expressed farnesyl-diphosphate synthase genes. J Biol Chem 271:7774–7780
Gaffe J, Bru JP, Causse M, Vidal A, Stamitti-Bert L, Carde JP, Gallusci P (2000) LEFPS1, a tomato farnesyl pyrophosphate gene highly expressed during early fruit development. Plant Physiol 123:1351–1362
Hemmerlin A, Rivera SB, Erickson HK, Poulter CD (2003) Enzymes encoded by the farnesyl diphosphate synthase gene family in the big sagebrush Artemisia tridentata ssp spiciformis. J Biol Chem 278:32132–32140
Cunillera N, Boronat A, Ferrer A (1997) The Arabidopsis thaliana FPS1 gene generates a novel mRNA that encodes a mitochondrial farnesyl-diphosphate synthase isoform. J Biol Chem 272:15381–15388
Thabet I, Guirimand G, Courdavault V, Papon N, Godet S, Dutilleul C, Bouzid S, Giglioli-Guivarc’h N, Clastre M, Simkin AJ (2011) The subcellular localization of periwinkle farnesyl diphosphate synthase provides insight into the role of peroxisome in isoprenoid biosynthesis. J Plant Physiol 168:2110–2116
Ito J, Batth TS, Petzold CJ, Redding-Johanson AM, Mukhopadhyay A, Verboom R, Meyer EH, Millar AH, Heazlewood JL (2011) Analysis of the Arabidopsis cytosolic proteome highlights subcellular partitioning of central plant metabolism. J Proteome Res 10:1571–1582
Reumann S, Quan S, Aung K, Yang P, Manandhar-Shrestha K, Holbrook D, Linka N, Switzenberg R, Wilkerson CG, Weber APM, Olsen LJ, Hu J (2009) In-depth proteome analysis of Arabidopsis leaf peroxisomes combined with in vivo subcellular targeting verification indicates novel metabolic and regulatory functions of peroxisomes. Plant Physiol 150:125–143
Closa M, Vranova E, Bortolotti C, Bigler L, Arro M, Ferrer A, Gruissem W (2010) The Arabidopsis thaliana FPP synthase isozymes have overlapping and specific functions in isoprenoid biosynthesis, and complete loss of FPP synthase activity causes early developmental arrest. Plant J 63:512–525
Keim V, Manzano D, Fernandez FJ, Closa M, Andrade P, Caudepon D, Bortolotti C, Vega MC, Arro M, Ferrer A (2012) Characterization of Arabidopsis FPS isozymes and FPS gene expression analysis provide insight into the biosynthesis of isoprenoid precursors in seeds. Plos One 7
Beck G, Coman D, Herren E, Ruiz-Sola M, Rodriguez-Concepcion M, Gruissem W, Vranova E (2013) Characterization of the GGPP synthase gene family in Arabidopsis thaliana. Plant Mol Biol 82:393–416
Ruppel NJ, Kropp KN, Davis PA, Martin AE, Luesse DR, Hangarter RP (2013) Mutations in geranylgeranyl diphosphate synthase 1 affect chloroplast development in Arabidopsis thaliana (Brassicaceae). Am J Bot 100:2074–2084
Dai ZB, Liu Y, Huang LQ, Zhang XL (2012) Production of miltiradiene by metabolically engineered Saccharomyces cerevisiae. Biotechnol Bioeng 109:2845–2853
Leonard E, Ajikumar PK, Thayer K, Xiao WH, Mo JD, Tidor B, Stephanopoulos G, Prather KLJ (2010) Combining metabolic and protein engineering of a terpenoid biosynthetic pathway for overproduction and selectivity control. Proc Natl Acad Sci USA 107:13654–13659
Vandermoten S, Haubruge E, Cusson M (2009) New insights into short-chain prenyltransferases: structural features, evolutionary history and potential for selective inhibition. Cell Mol Life Sci 66:3685–3695
Tarshis LC, Yan MJ, Poulter CD, Sacchettini JC (1994) Crystal structure of recombinant farnesyl diphosphate synthase at 2.6 Angstrom resolution. Biochemistry 33:10871–10877
Koyama T, Gotoh Y, Nishino T (2000) Intersubunit location of the active site of farnesyl diphosphate synthase: reconstruction of active enzymes by hybrid-type heteromeric dimers of site-directed mutants. Biochemistry 39:463–469
Hosfield DJ, Zhang YM, Dougan DR, Broun A, Tari LW, Swanson RV, Finn J (2004) Structural basis for bisphosphonate-mediated inhibition of isoprenoid biosynthesis. J Biol Chem 279:8526–8529
Chang TH, Guo RT, Ko TP, Wang AHJ, Liang PH (2006) Crystal structure of type-III geranylgeranyl pyrophosphate synthase from Saccharomyces cerevisiae and the mechanism of product chain length determination. J Biol Chem 281:14991–15000
Gabelli SB, McLellan JS, Montalvetti A, Oldfield E, Docampo R, Amzel LM (2006) Structure and mechanism of the farnesyl diphosphate synthase from Trypanosoma cruza: implications for drug design. Proteins Struct Funct Bioinform 62:80–88
Kavanagh KL, Dunford JE, Bunkoczi G, Russell RGG, Oppermann U (2006) The crystal structure of human geranylgeranyl pyrophosphate synthase reveals a novel hexameric arrangement and inhibitory product binding. J Biol Chem 281:22004–22012
Kloer DP, Welsch R, Beyer P, Schulz GE (2006) Structure and reaction geometry of geranylgeranyl diphosphate synthase from Sinapis alba. Biochemistry 45:15197–15204
Ohnuma S, Hirooka K, Hemmi H, Ishida C, Ohto C, Nishino T (1996) Conversion of product specificity of archaebacterial geranylgeranyl- diphosphate synthase. Identification of essential amino acid residues for chain length determination of prenyltransferase reaction. J Biol Chem 271:18831–18837
Ohnuma SI, Nakazawa T, Hemmi H, Hallberg AM, Koyama T, Ogura K, Nishino T (1996) Conversion from farnesyl diphosphate synthase to geranylgeranyl diphosphate synthase by random chemical mutagenesis. J Biol Chem 271:10087–10095
Wallrapp FH, Pan J-J, Ramamoorthy G, Almonacid DE, Hillerich BS, Seidel R, Patskovsky Y, Babbitt PC, Almo SC, Jacobson MP, Poulter CD (2013) Prediction of function for the polyprenyl transferase subgroup in the isoprenoid synthase superfamily. Proc Natl Acad Sci USA 110:E1196–E1202
Coman D, Altenhoff A, Zoller S, Gruissem W, Vranova E (2014) Distinct evolutionary strategies in the GGPPS family from plants. Front Plant Sci 5
Takahashi S, Koyama T (2006) Structure and function of cis-prenyl chain elongating enzymes. Chem Rec 6:194–205
Surmacz L, Swiezewska E (2011) Polyisoprenoids—secondary metabolites or physiologically important superlipids? Biochem Biophys Res Commun 407:627–632
Schmidt T, Lenders M, Hillebrand A, van Deenen N, Munt O, Reichelt R, Eisenreich W, Fischer R, Prufer D, Gronover CS (2010) Characterization of rubber particles and rubber chain elongation in Taraxacum koksaghyz. BMC Biochem 11
Sallaud C, Rontein D, Onillon S, Jabes F, Duffe P, Giacalone C, Thoraval S, Escoffier C, Herbette G, Leonhardt N, Causse M, Tissier A (2009) A novel pathway for sesquiterpene biosynthesis from Z,Z-farnesyl pyrophosphate in the wild tomato Solanum habrochaites. Plant Cell 21:301–317
Akhtar TA, Matsuba Y, Schauvinhold I, Yu G, Lees HA, Klein SE, Pichersky E (2013) The tomato cis-prenyltransferase gene family. Plant J 73:640–652
Bleeker PM, Mirabella R, Diergaarde PJ, VanDoorn A, Tissier A, Kant MR, Prins M, de Vos M, Haring MA, Schuurink RC (2012) Improved herbivore resistance in cultivated tomato with the sesquiterpene biosynthetic pathway from a wild relative. Proc Natl Acad Sci USA 109:20124–20129
Gonzales-Vigil E, Hufnagel DE, Kim J, Last RL, Barry CS (2012) Evolution of TPS20-related terpene synthases influences chemical diversity in the glandular trichomes of the wild tomato relative Solanum habrochaites. Plant J 71:921–935
Schilmiller AL, Schauvinhold I, Larson M, Xu R, Charbonneau AL, Schmidt A, Wilkerson C, Last RL, Pichersky E (2009) Monoterpenes in the glandular trichomes of tomato are synthesized from a neryl diphosphate precursor rather than geranyl diphosphate. Proc Natl Acad Sci USA 106:10865–10870
Gutensohn M, Nguyen TTH, McMahon RD III, Kaplan I, Pichersky E, Dudareva N (2014) Metabolic engineering of monoterpene biosynthesis in tomato fruits via introduction of the non-canonical substrate neryl diphosphate. Metab Eng 24:107–116
Demissie ZA, Erland LAE, Rheault MR, Mahmoud SS (2013) The biosynthetic origin of irregular monoterpenes in Lavandula: isolation and biochemical characterization of a novel cis-prenyl diphosphate synthase gene, lavendulyl diphosphate synthase. J Biol Chem 288:6333–6341
Surmacz L, Plochocka D, Kania M, Danikiewicz W, Swiezewska E (2014) cis-Prenyltransferase AtCPT6 produces a family of very short-chain polyisoprenoids in planta. Biochim Biophys Acta, Mol Cell Biol Lipids 1841:240–250
Kang J-H, Gonzales-Vigil E, Matsuba Y, Pichersky E, Barry CS (2014) Determination of residues responsible for substrate and product specificity of Solanum habrochaites short-chain cis-prenyltransferases. Plant Physiol 164:80–91
Kharel Y, Takahashi S, Yamashita S, Koyama T (2006) Manipulation of prenyl chain length determination mechanism of cis-prenyltransferases. FEBS J 273:647–657
Noike M, Katagiri T, Nakayama T, Koyama T, Nishino T, Hemmi H (2008) The product chain length determination mechanism of type II geranylgeranyl diphosphate synthase requires subunit interaction. FEBS J 275:3921–3933
Bohlmann J, Keeling CI (2008) Terpenoid biomaterials. Plant J 54:656–669
Chen F, Tholl D, Bohlmann J, Pichersky E (2011) The family of terpene synthases in plants: a mid-size family of genes for specialized metabolism that is highly diversified throughout the kingdom. Plant J 66:212–229
Degenhardt J, Kollner TG, Gershenzon J (2009) Monoterpene and sesquiterpene synthases and the origin of terpene skeletal diversity in plants. Phytochemistry 70:1621–1637
Tholl D, Chen F, Petri J, Gershenzon J, Pichersky E (2005) Two sesquiterpene synthases are responsible for the complex mixture of sesquiterpenes emitted from Arabidopsis flowers. Plant J 42:757–771
Chen F, Ro D-K, Petri J, Gershenzon J, Bohlmann J, Pichersky E, Tholl D (2004) Characterization of a root-specific Arabidopsis terpene synthase responsible for the formation of the volatile monoterpene 1,8-cineole. Plant Physiol 135:1956–1966
Kollner TG, Schnee C, Gershenzon J, Degenhardt J (2004) The sesquiterpene hydrocarbons of maize (Zea mays) form five groups with distinct developmental and organ-specific distribution. Phytochemistry 65:1895–1902
Xu MM, Wilderman PR, Peters RJ (2007) Following evolution’s lead to a single residue switch for diterpene synthase product outcome. Proc Natl Acad Sci USA 104:7397–7401
Keeling CI, Weisshaar S, Lin RPC, Bohlmann J (2008) Functional plasticity of paralogous diterpene synthases involved in conifer defense. Proc Natl Acad Sci USA 105:1085–1090
Greenhagen BT, O’Maille PE, Noel JP, Chappell J (2006) Identifying and manipulating structural determinates linking catalytic specificities in terpene synthases. Proc Natl Acad Sci USA 103:9826–9831
Davis EM, Croteau R (2000) Cyclization enzymes in the biosynthesis of monoterpenes, sesquiterpenes, and diterpenes. In: Leeper FJ, Vederas JC (eds) Topics in current chemistry: biosynthesis-aromatic polyketides, isoprenoids, alkaloids. Springer, Heidelberg, pp 53–95
Zi J, Mafu S, Peters RJ (2014) To gibberellins and beyond! Surveying the evolution of (di)terpenoid metabolism. Annu Rev Plant Biol 65:259–286
Hayashi K, Kawaide H, Notomi M, Sakigi Y, Matsuo A, Nozaki H (2006) Identification and functional analysis of bifunctional ent-kaurene synthase from the moss Physcomitrella patens. FEBS Lett 580:6175–6181
Anterola A, Shanle E, Mansouri K, Schuette S, Renzaglia K (2009) Gibberellin precursor is involved in spore germination in the moss Physcomitrella patens. Planta 229:1003–1007
Peters RJ, Carter OA, Zhang Y, Matthews BW, Croteau RB (2003) Bifunctional abietadiene synthase: mutual structural dependence of the active sites for protonation-initiated and ionization-initiated cyclizations. Biochemistry 42:2700–2707
Li GL, Kollner TG, Yin YB, Jiang YF, Chen H, Xu Y, Gershenzon J, Pichersky E, Chen F (2012) Nonseed plant Selaginella moellendorfii has both seed plant and microbial types of terpene synthases. Proc Natl Acad Sci USA 109:14711–14715
Geu-Flores F, Sherden NH, Courdavault V, Burlat V, Glenn WS, Wu C, Nims E, Cui Y, O’Connor SE (2012) An alternative route to cyclic terpenes by reductive cyclization in iridoid biosynthesis. Nature 492:138–142
Tundis R, Loizzo MR, Menichini F, Statti GA, Menichini F (2008) Biological and pharmacological activities of iridoids: recent developments. Mini-Rev Med Chem 8:399–420
Dewhirst SY, Pickett JA, Hardie J (2010) Aphid pheromones. In: Litwack G (ed) Vitamins and hormones: pheromones, pp 551–574
Koksal M, Zimmer I, Schnitzler JP, Christianson DW (2010) Structure of isoprene synthase illuminates the chemical mechanism of teragram atmospheric carbon emission. J Mol Biol 402:363–373
Whittington DA, Wise ML, Croteau R, Christianson DW (2002) Insights into monoterpene cyclization reactions in biology: crystal structure of (+)-bornyl diphosphate synthase. Biochemistry 41:8973
Whittington DA, Wise ML, Urbansky M, Coates RM, Croteau RB, Christianson DW (2002) Bornyl diphosphate synthase: structure and strategy for carbocation manipulation by a terpenoid cyclase. Proc Natl Acad Sci USA 99:15375–15380
Hyatt DC, Youn BY, Zhao YX, Santhamma B, Coates RM, Croteau RB, Kang CH (2007) Structure of limonene synthase, a simple model for terpenoid cyclase catalysis. Proc Natl Acad Sci USA 104:5360–5365
Kampranis SC, Ioannidis D, Purvis A, Mahrez W, Ninga E, Katerelos NA, Anssour S, Dunwell JM, Degenhardt J, Makris AM, Goodenough PW, Johnson CB (2007) Rational conversion of substrate and product specificity in a Salvia monoterpene synthase: structural insights into the evolution of terpene synthase function. Plant Cell 19:1994–2005
Starks CM, Back KW, Chappell J, Noel JP (1997) Structural basis for cyclic terpene biosynthesis by tobacco 5-epi-aristolochene synthase. Science 277:1815–1820
Gennadios HA, Gonzalez V, Di Costanzo L, Li AA, Yu FL, Miller DJ, Allemann RK, Christianson DW (2009) Crystal structure of (+)-delta-cadinene synthase from Gossypium arboreum and evolutionary divergence of metal binding motifs for catalysis. Biochemistry 48:6175–6183
Koksal M, Jin YH, Coates RM, Croteau R, Christianson DW (2011) Taxadiene synthase structure and evolution of modular architecture in terpene biosynthesis. Nature 469:116–120
Koksal M, Potter K, Peters RJ, Christianson DW (2014) 1.55 angstrom-resolution structure of ent-copalyl diphosphate synthase and exploration of general acid function by site-directed mutagenesis. Biochim Biophys Acta, Gen Sub 1840:184–190
Koeksal M, Hu H, Coates RM, Peters RJ, Christianson DW (2011) Structure and mechanism of the diterpene cyclase ent-copalyl diphosphate synthase. Nat Chem Biol 7:431–433
Zhou K, Gao Y, Hoy JA, Mann FM, Honzatko RB, Peters RJ (2012) Insights into diterpene cyclization from structure of bifunctional abietadiene synthase from Abies grandis. J Biol Chem 287:6840–6850
Cao R, Zhang YH, Mann FM, Huang CC, Mukkamala D, Hudock MP, Mead ME, Prisic S, Wang K, Lin FY, Chang TK, Peters RJ, Odfield E (2010) Diterpene cyclases and the nature of the isoprene fold. Proteins: Struct Funct Bioinform 78:2417–2432
Gao Y, Honzatko RB, Peters RJ (2012) Terpenoid synthase structures: a so far incomplete view of complex catalysis. Nat Prod Rep 29:1153–1175
Christianson DW (2006) Structural biology and chemistry of the terpenoid cyclases. Chem Rev 106:3412–3442
Aubourg S, Lecharny A, Bohlmann J (2002) Genomic analysis of the terpenoid synthase (AtTPS) gene family of Arabidopsis thaliana. Mol Genet Genomics 267:730–745
Tholl D, Lee S (2011) Terpene specialized metabolism in Arabidopsis thaliana. The Arabidopsis Book 9:e0143
Field B, Osbourn AE (2008) Metabolic diversification-independent assembly of operon-like gene clusters in different plants. Science 320:543–547
Field B, Fiston-Lavier AS, Kemen A, Geisler K, Quesneville H, Osbourn AE (2011) Formation of plant metabolic gene clusters within dynamic chromosomal regions. Proc Natl Acad Sci USA 108:16116–16121
Mugford ST, Louveau T, Melton R, Qi XQ, Bakht S, Hill L, Tsurushima T, Honkanen S, Rosser SJ, Lomonossoff GP, Osbourn A (2013) Modularity of plant metabolic gene clusters: a trio of linked genes that are collectively required for acylation of triterpenes in oat. Plant Cell 25:1078–1092
Wilderman PR, Xu MM, Jin YH, Coates RM, Peters RJ (2004) Identification of syn-pimara-7,15-diene synthase reveals functional clustering of terpene synthases involved in rice phytoalexin/allelochemical biosynthesis. Plant Physiol 135:2098–2105
Falara V, Akhtar TA, Nguyen TTH, Spyropoulou EA, Bleeker PM, Schauvinhold I, Matsuba Y, Bonini ME, Schilmiller AL, Last RL, Schuurink RC, Pichersky E (2011) The tomato terpene synthase gene family. Plant Physiol 157:770–789
Segura MJR, Jackson BE, Matsuda SPT (2003) Mutagenesis approaches to deduce structure-function relationships in terpene synthases. Nat Prod Rep 20:304–317
Phillips DR, Rasbery JM, Bartel B, Matsuda SPT (2006) Biosynthetic diversity in plant triterpene cyclization. Curr Opin Plant Biol 9:305–314
Wegel E, Koumproglou R, Shaw P, Osbourn A (2009) Cell type-specific chromatin decondensation of a metabolic gene cluster in oats. Plant Cell 21:3926–3936
Mylona P, Owatworakit A, Papadopoulou K, Jenner H, Qin B, Findlay K, Hill L, Qi X, Bakht S, Melton R, Osbourn A (2008) Sad3 and Sad4 are required for saponin biosynthesis and root development in oat. Plant Cell 20:201–212
Yamane H (2013) Biosynthesis of phytoalexins and regulatory mechanisms of it in rice. Biosci Biotech Biochem 77:1141–1148
Xu YH, Wang JW, Wang S, Wang JY, Chen XY (2004) Characterization of GaWRKY1, a cotton transcription factor that regulates the sesquiterpene synthase gene (+)-delta-cadinene synthase-A. Plant Physiol 135:507–515
Lu X, Zhang L, Zhang FY, Jiang WM, Shen Q, Zhang LD, Lv ZY, Wang GF, Tang KX (2013) AaORA, a trichome-specific AP2/ERF transcription factor of Artemisia annua, is a positive regulator in the artemisinin biosynthetic pathway and in disease resistance to Botrytis cinerea. New Phytol 198:1191–1202
Reeves PH, Ellis CM, Ploense SE, Wu MF, Yadav V, Tholl D, Chetelat A, Haupt I, Kennerley BJ, Hodgens C, Farmer EE, Nagpal P, Reed JW (2012) A regulatory network for coordinated flower maturation. PLoS Genet 8
Hong GJ, Xue XY, Mao YB, Wang LJ, Chen XY (2012) Arabidopsis MYC2 interacts with DELLA proteins in regulating sesquiterpene synthase gene expression. Plant Cell 24:2635–2648
Dombrecht B, Xue GP, Sprague SJ, Kirkegaard JA, Ross JJ, Reid JB, Fitt GP, Sewelam N, Schenk PM, Manners JM, Kazan K (2007) MYC2 differentially modulates diverse jasmonate-dependent functions in Arabidopsis. Plant Cell 19:2225–2245
Yadav V, Mallappa C, Gangappa SN, Bhatia S, Chattopadhyay S (2005) A basic helix-loop-helix transcription factor in Arabidopsis, MYC2, acts as a repressor of blue light-mediated photomorphogenic growth. Plant Cell 17:1953–1966
Vaughan MM, Wang Q, Webster FX, Kiemle D, Hong YJ, Tantillo DJ, Coates RM, Wray AT, Askew W, O’Donnell C, Tokuhisa JG, Tholl D (2013) Formation of the unusual semivolatile diterpene rhizathalene by the Arabidopsis class I terpene synthase TPS08 in the root stele is involved in defense against belowground herbivory. Plant Cell 25:1108–1125
Ro DK, Ehlting J, Keeling CI, Lin R, Mattheus N, Bohlmann J (2006) Microarray expression profiling and functional characterization of AtTPS genes: duplicated Arabidopsis thaliana sesquiterpene synthase genes At4g13280 and At4g13300 encode root-specific and wound-inducible (Z)-γ-bisabolene synthases. Arch Biochem Biophys 448:104–116
Loreto F, Dicke M, Schnitzler JP, Turlings TCJ (2014) Plant volatiles and the environment. Plant Cell Environ 37:1905–1908
Behnke K, Ehlting B, Teuber M, Bauerfeind M, Louis S, Hasch R, Polle A, Bohlmann J, Schnitzler JP (2007) Transgenic, non-isoprene emitting poplars don’t like it hot. Plant J 51:485–499
Sharkey TD, Yeh SS (2001) Isoprene emission from plants. Annu Rev Plant Physiol Plant Mol Biol 52:407–436
Velikova V, Ghirardo A, Vanzo E, Merl J, Hauck SM, Schnitzler J-P (2014) Genetic manipulation of isoprene emissions in poplar plants remodels the chloroplast proteome. J Proteome Res 13:2005–2018
Behnke K, Kleist E, Uerlings R, Wildt J, Rennenberg H, Schnitzler JP (2009) RNAi-mediated suppression of isoprene biosynthesis in hybrid poplar impacts ozone tolerance. Tree Physiol 29:725–736
Loreto F, Schnitzler JP (2010) Abiotic stresses and induced BVOCs. Trends Plant Sci 15:154–166
Vickers CE, Possell M, Laothawornkitkul J, Ryan AC, Hewitt CN, Mullineaux PM (2011) Isoprene synthesis in plants: lessons from a transgenic tobacco model. Plant Cell Environ 34:1043–1053
Schnitzler JP, Louis S, Behnke K, Loivamaki M (2010) Poplar volatiles—biosynthesis, regulation and (eco)physiology of isoprene and stress-induced isoprenoids. Plant Biol 12:302–316
Byers K, Bradshaw HD, Riffell JA (2014) Three floral volatiles contribute to differential pollinator attraction in monkeyflowers (Mimulus). J Exp Biol 217:614–623
Huang M, Sanchez-Moreiras AM, Abel C, Sohrabi R, Lee S, Gershenzon J, Tholl D (2012) The major volatile organic compound emitted from Arabidopsis thaliana flowers, the sesquiterpene (E)-β-caryophyllene, is a defense against a bacterial pathogen. New Phytol 193:997–1008
Junker RR, Loewel C, Gross R, Dötterl S, Keller A, Blüthgen N (2011) Composition of epiphytic bacterial communities differs on petals and leaves. Plant Biol 13:918–924
Wang H, Guo WF, Zhang PJ, Wu ZY, Liu SS (2008) Experience-induced habituation and preference towards non-host plant odors in ovipositing females of a moth. J Chem Ecol 34:330–338
Laothawornkitkul J, Paul ND, Vickers CE, Possell M, Taylor JE, Mullineaux PM, Hewitt CN (2008) Isoprene emissions influence herbivore feeding decisions. Plant Cell Environ 31:1410–1415
Bleeker PM, Diergaarde PJ, Ament K, Schutz S, Johne B, Dijkink J, Hiemstra H, de Gelder R, de Both MTJ, Sabelis MW, Haring MA, Schuurink RC (2011) Tomato-produced 7-epizingiberene and R-curcumene act as repellents to whiteflies. Phytochemistry 72:68–73
Zulak KG, Bohlmann J (2010) Terpenoid biosynthesis and specialized vascular cells of conifer defense. J Integr Plant Biol 52:86–97
Hall DE, Robert JA, Keeling CI, Domanski D, Quesada AL, Jancsik S, Kuzyk MA, Hamberger B, Borchers CH, Bohlmann J (2011) An integrated genomic, proteomic and biochemical analysis of (+)-3-carene biosynthesis in Sitka spruce (Picea sitchensis) genotypes that are resistant or susceptible to white pine weevil. Plant J 65:936–948
Gols R (2014) Direct and indirect chemical defences against insects in a multitrophic framework. Plant Cell Environ 37:1741–1752
Pierik R, Ballare CL, Dicke M (2014) Ecology of plant volatiles: taking a plant community perspective. Plant Cell Environ 37:1845–1853
Schnee C, Kollner TG, Held M, Turlings TCJ, Gershenzon J, Degenhardt J (2006) The products of a single maize sesquiterpene synthase form a volatile defense signal that attracts natural enemies of maize herbivores. Proc Natl Acad Sci USA 103:1129–1134
Fontana A, Held M, Fantaye CA, Turlings TC, Degenhardt J, Gershenzon J (2011) Attractiveness of constitutive and herbivore-induced sesquiterpene blends of maize to the parasitic wasp Cotesia marginiventris (Cresson). J Chem Ecol 37:582–591
Kappers IF, Aharoni A, van Herpen TWJM, Luckerhoff LLP, Dicke M, Bouwmeester HJ (2005) Genetic engineering of terpenoid metabolism attracts bodyguards to Arabidopsis. Science 309:2070–2072
McCormick AC, Unsicker SB, Gershenzon J (2012) The specificity of herbivore-induced plant volatiles in attracting herbivore enemies. Trends Plant Sci 17:303–310
Kessler A, Heil M (2011) The multiple faces of indirect defences and their agents of natural selection. Funct Ecol 25:348–357
Hilker M, Meiners T (2006) Early herbivore alert: insect eggs induce plant defense. J Chem Ecol 32:1379–1397
Buchel K, Malskies S, Mayer M, Fenning TM, Gershenzon J, Hilker M, Meiners T (2011) How plants give early herbivore alert: volatile terpenoids attract parasitoids to egg-infested elms. Basic Appl Ecol 12:403–412
Arimura G, Ozawa R, Shimoda T, Nishioka T, Boland W, Takabayashi J (2000) Herbivory-induced volatiles elicit defence genes in lima bean leaves. Nature 406:512–515
Frost CJ, Appel M, Carlson JE, De Moraes CM, Mescher MC, Schultz JC (2007) Within-plant signalling via volatiles overcomes vascular constraints on systemic signalling and primes responses against herbivores. Ecol Lett 10:490–498
Heil M, Karban R (2010) Explaining evolution of plant communication by airborne signals. Trends Ecol Evol 25:137–144
Heil M (2014) Herbivore-induced plant volatiles: targets, perception and unanswered questions. New Phytol. doi:10.1111/nph.12977
Himanen SJ, Blande JD, Klemola T, Pulkkinen J, Heijari J, Holopainen JK (2010) Birch (Betula spp.) leaves adsorb and re-release volatiles specific to neighbouring plants—a mechanism for associational herbivore resistance? New Phytol 186:722–732
Runyon JB, Mescher MC, De Moraes CM (2006) Volatile chemical cues guide host location and host selection by parasitic plants. Science 313:1964–1967
Jassbi AR, Zamanizadehnajari S, Baldwin IT (2010) 17-Hydroxygeranyllinalool glycosides are major resistance traits of Nicotiana obtusifolia against attack from tobacco hornworm larvae. Phytochemistry 71:1115–1121
Schmelz EA, Kaplan F, Huffaker A, Dafoe NJ, Vaughan MM, Ni XZ, Rocca JR, Alborn HT, Teal PE (2011) Identity, regulation, and activity of inducible diterpenoid phytoalexins in maize. Proc Natl Acad Sci USA 108:5455–5460
Huffaker A, Kaplan F, Vaughan MM, Dafoe NJ, Ni XZ, Rocca JR, Alborn HT, Teal PEA, Schmelz EA (2011) Novel acidic sesquiterpenoids constitute a dominant class of pathogen-induced phytoalexins in maize. Plant Physiol 156:2082–2097
Kuzina V, Ekstrom CT, Andersen SB, Nielsen JK, Olsen CE, Bak S (2009) Identification of defense compounds in Barbarea vulgaris against the herbivore Phyllotreta nemorum by an ecometabolomic approach. Plant Physiol 151:1977–1990
Rasmann S, Kollner TG, Degenhardt J, Hiltpold I, Toepfer S, Kuhlmann U, Gershenzon J, Turlings TCJ (2005) Recruitment of entomopathogenic nematodes by insect-damaged maize roots. Nature 434:732–737
Turlings TCJ, Hiltpold I, Rasmann S (2012) The importance of root-produced volatiles as foraging cues for entomopathogenic nematodes. Plant Soil 358:47–56
Degenhardt J, Hiltpold I, Kollner TG, Frey M, Gierl A, Gershenzon J, Hibbard BE, Ellersieck MR, Turlings TCJ (2009) Restoring a maize root signal that attracts insect-killing nematodes to control a major pest. Proc Natl Acad Sci USA 106:13213–13218
Robert CAM, Erb M, Hiltpold I, Hibbard BE, Gaillard MDP, Bilat J, Degenhardt J, Cambet-Petit-Jean X, Turlings TCJ, Zwahlen C (2013) Genetically engineered maize plants reveal distinct costs and benefits of constitutive volatile emissions in the field. Plant Biotechnol J 11:628–639
Xu MM, Galhano R, Wiemann P, Bueno E, Tiernan M, Wu W, Chung IM, Gershenzon J, Tudzynski B, Sesma A, Peters RJ (2012) Genetic evidence for natural product-mediated plant–plant allelopathy in rice (Oryza sativa). New Phytol 193:570–575
Thimmappa R, Geisler K, Louveau T, O’Maille P, Osbourn A (2014) Triterpene biosynthesis in plants. Annu Rev Plant Biol 65:225–257
Kemen AC, Honkanen S, Melton RE, Findlay KC, Mugford ST, Hayashi K, Haralampidis K, Rosser SJ, Osbourn A (2014) Investigation of triterpene synthesis and regulation in oats reveals a role for beta-amyrin in determining root epidermal cell patterning. Proc Natl Acad Sci USA 111:8679–8684
Chaturvedi R, Venables B, Petros RA, Nalam V, Li MY, Wang XM, Takemoto LJ, Shah J (2012) An abietane diterpenoid is a potent activator of systemic acquired resistance. Plant J 71:161–172
Waldie T, McCulloch H, Leyser O (2014) Strigolactones and the control of plant development: lessons from shoot branching. Plant J 79:607–622
Akiyama K, Matsuzaki K, Hayashi H (2005) Plant sesquiterpenes induce hyphal branching in arbuscular mycorrhizal fungi. Nature 435:824–827
Matusova R, Rani K, Verstappen FWA, Franssen MCR, Beale MH, Bouwmeester HJ (2005) The strigolactone germination stimulants of the plant-parasitic Striga and Orobanche spp. are derived from the carotenoid pathway. Plant Physiol 139:920–934
Gomez-Roldan V, Fermas S, Brewer PB, Puech-Pages V, Dun EA, Pillot JP, Letisse F, Matusova R, Danoun S, Portais JC, Bouwmeester H, Becard G, Beveridge CA, Rameau C, Rochange SF (2008) Strigolactone inhibition of shoot branching. Nature 455:189–U122
Domagalska MA, Leyser O (2011) Signal integration in the control of shoot branching. Nat Rev Mol Cell Biol 12:211–221
Brewer PB, Koltai H, Beveridge CA (2013) Diverse roles of strigolactones in plant development. Mol Plant 6:18–28
Rodriguez S, Kirby J, Denby CM, Keasling JD (2014) Production and quantification of sesquiterpenes in Saccharomyces cerevisiae, including extraction, detection and quantification of terpene products and key related metabolites. Nat Protoc 9:1980–1996
Acknowledgments
This work was supported by the National Science Foundation MCB grant 0950865.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Tholl, D. (2015). Biosynthesis and Biological Functions of Terpenoids in Plants. In: Schrader, J., Bohlmann, J. (eds) Biotechnology of Isoprenoids. Advances in Biochemical Engineering/Biotechnology, vol 148. Springer, Cham. https://doi.org/10.1007/10_2014_295
Download citation
DOI: https://doi.org/10.1007/10_2014_295
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-20106-1
Online ISBN: 978-3-319-20107-8
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)